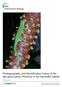
Natural hybridization in lizards of the genus Tupinambis (Teiidae) in the southernmost contact zone of their distribution range
|
|
|
- Louisa Wiggins
- 6 years ago
- Views:
Transcription
1 Ann. Zool. Fennici 51: ISSN X (print), ISSN (online) Helsinki 30 June 2014 Finnish Zoological and Botanical Publishing Board 2014 Natural hybridization in lizards of the genus Tupinambis (Teiidae) in the southernmost contact zone of their distribution range Imanol Cabaña 1,2, Cristina N. Gardenal 1, Margarita Chiaraviglio 2 & Paula C. Rivera 1,2, * 1) Laboratorio de Genética de Poblaciones y Evolución, Instituto de Diversidad y Ecología Animal (CONICET-UNC) and Facultad de Ciencias Exactas, Físicas y Naturales, Universidad Nacional de Córdoba, Av. Vélez Sarsfield 299, Córdoba, Argentina (*corresponding author s paularivera1@gmail.com) 2) Laboratorio de Biología del Comportamiento, Instituto de Diversidad y Ecología Animal (CONICET-UNC) and Facultad de Ciencias Exactas, Físicas y Naturales, Universidad Nacional de Córdoba, Av. Vélez Sarsfield 299, Córdoba, Argentina Received 20 May 2013, final version received 15 July 2013, accepted 19 Aug Cabaña, I., Gardenal, C. N., Chiaraviglio, M. & Rivera, P. C. 2014: Natural hybridization in lizards of the genus Tupinambis (Teiidae) in the southernmost contact zone of their distribution range. Ann. Zool. Fennici 51: Studies on the mechanisms of speciation and maintenance of lineages have paid great attention to hybridization between species because this process is considered an important source of variability and evolution. In recent years, the use of molecular markers has provided more detailed information on the distribution and magnitude of hybridization in natural populations. Here we present a phylogenetic analysis using one mitochondrial and one nuclear DNA segment as molecular markers in two closely related lizard species, Tupinambis merianae and T. rufescens, which are present in a continuous area including allopatric and sympatric populations. Consensus trees obtained with the mitochondrial gene showed two well-supported clades. Some individuals clustered with one of the species in the tree obtained with mitochondrial DNA, and with the other species in the tree recovered using the nuclear gene, demonstrating the occurrence of hybridization between these species. Hybrid individuals were captured in the area of sympatry, suggesting the existence of a hybrid zone in the contact area of the distribution ranges of these two lizards, which corresponds to the ecotone between Dry Chaco and Espinal. This work presents the first evidence of natural hybridization within the genus Tupinambis. Introduction Hybridization between species is an interesting process in speciation and lineage maintenance. By backcrossing, hybridization may result in the transfer of alleles between species; the introgression of new genetic information can be an important source of variability and subsequent evolution, even at low levels (Anderson 1949, Barton 2001, Seehausen 2004, Mallet 2007, 2008).
2 Ann. Zool. Fennici Vol. 51 Hybridization in Tupinambis 341 Hybrid individuals may originate from mating between females of one species and males of the other species and vice versa (reciprocal hybridization), or from mating of females of one species with males of the other species only (unidirectional hybridization) (Wirtz 1999). Detection of hybrid individuals has traditionally been performed by morphological, cytogenetic and/or histocompatibility studies (Dowling & Secor 1997). Today, the use of molecular markers enables the study of directionality, distribution and/or extent of hybridization in natural populations (Mallet 2008). Natural hybridization has been studied in teiids of the genera Aspidoscelis, Cnemidophorus and Kentropix (Dessauer et al. 2000, Taylor et al. 2001, 2003, Reeder et al. 2002, Cole et al. 2007, Manríquez-Morán 2007) with the aim to identify the parental species that gave rise to parthenogenetic forms. None of these studies included hybrids able to sexually reproduce among them or with their parental species. To identify the origin of hybrids and obtain a comprehensive picture of lineage evolution, it is convenient to combine analysis with mitochondrial (mtdna) and nuclear (ndna) DNA markers, since the ndna evolves at lower rates than the mtdna, recombines and represents the history of both parents, rather than that of the maternal lineage (Leache & McGuire 2006, Zarza et al. 2008). If hybrids share mtdna haplotypes with only one of the parental species, the process behind it is unidirectional hybridization. However, if some hybrids share haplotypes with one of the parental species and other hybrids do so with the other parental species, reciprocal hybridization can be assumed. The genus Tupinambis belongs to the family Teiidae, with their two southernmost distributed species being present in Argentina. Tupinambis merianae is widespread, spanning several biogeographic regions, whereas T. rufescens has a range restricted mainly to Arid Chaco (Cei 1986, 1993). Although the habitats of these species differ, in central and north-central Argentina their distributions overlap in the ecotones of Arid Chaco and Espinal, and Arid Chaco and Humid Chaco. Hitherto, there is no evidence of hybridization in natural populations of the genus Tupinambis. Cei (1993) proposed the existence of a possible hybrid specimen between T. merianae and T. rufescens on the basis of its pattern scalation. Fitzgerald et al. (1999) found a paraphyletic relationship between T. rufescens and T. duseni and argued incomplete lineage sorting as the most probable explanation for their results, introgression of mtdna being a less likely possibility since the species are not in sympatry. In the present study, we evaluate the occurrence of hybridization between T. merianae and T. rufescens in the southernmost contact zone of their distribution, by analyzing ndna and mtdna data using a phylogenetic approach. We also explore the directionality and distribution of this phenomenon. Material and methods Study area and data collection The sample area was located in central Argentina (29 32 S and W; Fig. 1), and covered two biogeographic regions, Arid Chaco and Espinal, and their ecotone zone. Individuals belonging to T. merianae and T. rufescens were identified phenotypically on the basis of their coloration according to Cei (1993), who established coloration as a valid character to differentiate these species: T. merianae is dark olive green, sometimes almost black, and T. rufescens is reddish. These individuals were from localities belonging to areas of sympatry and allopatry of those species, as determined in Cardozo et al. (2012). We obtained samples (muscle tissue or scales) from individuals hunted by rural people, since commercial exploitation is permitted (Porini 2006), and from road-killed individuals. In both cases, we recorded the coordinates using GPS. All tissue samples were stored in 70% ethanol at 20 C and were deposited in the tissue collection of the Behavioural Biology Laboratory (IDEA, CONICET-UNC). Scientific capture was authorized by the government environmental agency. Genomic DNA extraction and sequencing Genomic DNA was obtained from muscle tissue
3 342 Cabaña et al. Ann. ZOOL. Fennici Vol. 51 A 29 S 65 W 61 W 32 S km B 65 W 61 W 29 S 32 S km Fig. 1. Spatial distribution of localities, sample size and frequency of (A) ND4 haplotypes and (B) ACA4 alleles. The dots show the localities (black dots: populations of T. rufescens in allopatry; light-gray dots: populations of T. merianae in allopatry; dark-gray dots: populations in the sympatric area). The circles represent the sample size for each species at each locality (black slices: individuals of T. rufescens; light-gray slices: individuals of T. merianae; white slices: hybrid individuals classified as T. merianae with mtdna from T. rufescens; dark gray slice: hybrid individual classified as T. rufescens with mtdna from T. merianae). Letters within the circles indicated haplotypes for ND4 gene in A, and alleles for ACA4 gene in B. For explanations see Table 1. or scales, using a saline extraction method (Bruford et al. 1992). We used a fragment of the Nicotinamide Adenine Dinucleotide Dehydrogenase subunit 4 gene (ND4) as the mtdna marker and a fragment of the α-cardiac-actin Intron 4 gene (ACA4) as the ndna marker. Both markers showed high polymorphism and have been used in previous studies on Squamata (Pinho et al. 2007, Giffor & Larson 2008, respectively). We performed amplification of the ND4 and ACA4 genes via the polymerase chain reaction (PCR) using specific primers described by Forstner et al. (1995) and Waltari and Edwards (2002), respectively, following the protocol of Martínez et al. (2009) with an annealing temperature of 50 C. All purifications and sequencing reactions were performed by Macrogen Inc. (Seoul, South Korea) in 3 to 5 direction. The sequences obtained in the present study were deposited in GenBank (Table 1).
4 Ann. Zool. Fennici Vol. 51 Hybridization in Tupinambis 343 Phylogenetic analysis Chromatograms were examined using Chromas lite ver (Technelysium Pty Ltd., USA). Sequences were aligned using the Muscle software (Edgar 2004) with the default parameters. In the case of ND4, sequences were translated into amino acids to confirm alignment. For the nuclear gene, alleles were determined using the PHASE software (Stephens & Donnelly 2003), which allowed us to estimate the allele that was more likely to occur when a sequence had more than one heterozygous site, by using the probabilistic approach. This method assumes Hardy- Weinberg equilibrium and uses a coalescentbased Bayesian method to infer haplotypes. The file input used in this software was obtained from SeqPHASE (available at fr/seqphase/). The number of iterations, thinning intervals and burn-in values were the default parameters. We estimated phylogenetic relationships using two approaches, Maximum Parsimony (MP) and Bayesian inference (BI), for mtdna (ND4) and ndna (ACA4), separately. For MP analysis, we used PAUP 4.0b10 (Swofford 1998) software considering equal weighting for all characters. The node support was evaluated with 1000 bootstrap replicates. For BI, we estimated HKY + G for the mitochondrial gene and F81 for the nuclear gene as the most appropriate models of sequence evolution using JModeltest (Posada 2008), under the Akaike information criterion. Bayesian analyses were performed using MrBayes (Ronquist & Huelsenbeck 2003). For each data set, the analyses were made for two million generations. In both analyses two independent runs were simultaneously performed on the data, each using one cold and three heated chains, with sampling intervals of 1000 generations. We discarded the first 25% of the samples as burn-in. We determined support for tree nodes according to the values of Bayesian posterior probability obtained from a majority-rule consensus tree. We included sequences of the ND4 gene for T. merianae and T. rufescens available from GenBank. Based on previous phylogenies (Fitzgerald et al. 1999, Giugliano et al. 2007) and sequence availability, we used T. quadrilineatus and T. longilineus as outgroups for phylogenetic reconstructions with the ND4 gene and Ameiva chrysolaema with the ACA4 gene. To get a clear picture of the haplotype and allele frequencies, and the relationships among the co-existing lineages, we constructed networks for each gene. They were obtained with a median-joining approach using the program Network (Bandelt et al. 1999) with default parameters. For ACA4, we considered both alleles for each individual. Results We analyzed 29 individuals phenotypically classified as T. merianae from eight localities and 19 individuals classified as T. rufescens from six localities (Table 1 and Fig. 1). An 807-bp fragment of the mtdna gene and a 413-bp fragment of the ndna gene were sequenced from each specimen. All the different sequences obtained were deposited in GenBank (accession numbers KF KF034101). The MP and BI phylogenetic trees using the ND4 data matrix showed great congruency and similar topology in estimating phylogenetic relationships among the taxa (Fig. 2A). Two well supported clades were obtained, one with most of the T. merianae sequences and the other one with most of the T. rufescens sequences. Four individuals identified phenotypically as T. merianae presented sequences grouped within the clade corresponding to T. rufescens and one individual classified as T. rufescens presented an ND4 haplotype grouped within the T. merianae clade. The MP and BI phylogenetic trees using the ACA4 data matrix also showed similar topology in estimating phylogenetic relationships among the taxa (Fig. 2B). Sequences of T. merianae form a polytomy that also includes all specimens classified as T. merianae but presenting T. rufescens mtdna. Only one well supported clade grouped sequences of T. rufescens. The sequence of the individual identified phenotypically as T. rufescens and presenting T. merianae mtdna was grouped within this clade, with high posterior probability in the Bayesian analysis. In summary, five individuals had nuclear alleles of one species, according to their pheno-
5 344 Cabaña et al. Ann. ZOOL. Fennici Vol. 51 Table 1. Localities, coordinates, specimens captured and GenBank accession numbers for each haplotype for ND4 gene or allele for ACA4 gene sequence for the two species of Tupinambis (the letters in parentheses next to the numbers are codes used in Figs. 1 and 3). Individuals were classified according to skin coloration (Cei 1993). Individual identification that starts with R stands for T. rufescens and with M, for T. merianae. Hybrid individuals are denoted with an asterisk (*). Locality Latitude Longitude Individual GenBank accession numbers ND4 ACA4 ACA R490 KF (H) KF (R) KF (R) R491 KF (I) KF (R) KF (R) R590 KF (E) KF (R) KF (R) R601 KF (I) KF (R) KF (R) R606 KF (G) KF (R) KF (R) R607 KF (E) KF (R) KF (R) R610 KF (G) KF (R) KF (R) R586 KF (I) KF (R) KF (R) R598 KF (F) KF (Q) KF (R) R608 KF (F) KF (R) KF (R) R587 KF (I) KF (R) KF (R) R734 KF (I) KF (R) KF (R) R735 KF (I) KF (R) KF (R) R722 KF (I) KF (P) KF (R) R723 KF (H) KF (R) KF (R) R741 KF (J) KF (R) KF (R) RIC23 KF (J) KF (R) KF (R) MIC28 KF (A) KF (L) KF (L) R681* KF (A) KF (R) KF (R) M691* KF (J) KF (M) KF (O) M694 KF (A) KF (L) KF (L) M697 KF (A) KF (L) KF (N) R744 KF (H) KF (R) KF (R) MIC26 KF (A) KF (L) KF (L) MIC27 KF (A) KF (K) KF (K) MIC31* KF (J) KF (L) KF (L) MIC38 KF (A) KF (K) KF (O) MIC6* KF (I) KF (M) KF (M) MIC8 KF (A) KF (O) KF (O) MIC17 KF (A) KF (K) KF (M) M507 KF (A) KF (L) KF (K) M54 KF (A) KF (M) KF (M) M55 KF (A) KF (M) KF (M) M56 KF (A) KF (L) KF (M) M57 KF (A) KF (L) KF (M) M58 KF (A) KF (L) KF (O) M427 KF (A) KF (M) KF (M) M341 KF (A) KF (L) KF (M) M342* KF (J) KF (L) KF (M) M303 KF (B) KF (L) KF (M) M304 KF (D) KF (L) KF (O) M305 KF (B) KF (K) KF (O) M306 KF (A) KF (L) KF (M) M391 KF (C) KF (L) KF (O) M392 KF (A) KF (L) KF (O) MIC47 KF (C) KF (L) KF (K) MIC48 KF (B) KF (L) KF (N) MIC49 KF (C) KF (M) KF (O)
6 Ann. Zool. Fennici Vol. 51 Hybridization in Tupinambis 345 A AF T. quadrilineatus 0.97/100 Af AF T. longilineus 0.66/ / /66 AF AF KF (5*, 6, 7, 8, 9, 10, 11) AF KF (11, 12) KF (11, 12) T. merianae 1.00/ /87 AF KF (11) KF (1) KF (1, 2, 3, 4, 8*) KF (1, 4, 6) B 0.1 KF (6, 12) 0.96/100 KF (2) T. rufescens AF AF /87 KF (1) 1.00/82 KF (5,6*, 7*, 10*) A. chrysolaema KF (5, 6, 7*, 8, 9, 10*, 11, 12) KF (7, 8, 11, 12) KF (6*, 8*, 9, 10*, 11, 12) T. merianae KF (6*, 7, 8, 9, 11, 12) 0.58/ KF (4) 1.00/94 KF (1, 2, 3, 4, 5*, 6) KF (2) 0.1 T. rufescens Fig. 2. Consensus tree obtained from Bayesian inference with the matrix of (A) mtdna (ND4) and (B) ndna (ACA4). Node support has the following order: Bayesian posterior probability/mp after 1000 bootstrap replicates. The localities (numbers as in Table 1) where each haplotype or allele was found are given in parentheses; the localities where hybrid individuals were found are indicated with an asterisk (*). typic classification, but showed mitochondrial sequences corresponding to the other species, i.e., they presented introgressed haplotypes. All these specimens were located in the areas of sympatry of both species (localities 5, 6, 7, 8 and 10) (Table 1 and Fig. 1A). Our results showed four haplotypes for the ND4 gene and five alleles for the ACA4 gene in T. merianae, whereas in T. rufescens, six haplotypes were identified for the mtdna gene and three alleles for the ndna gene (Fig. 2). Introgressed haplotypes were not considered when counting the haplotypes of each species. The haplotype networks obtained showed two well-separated groups (Fig. 3), one belonging to T. merianae and the other to T. rufescens.
7 346 Cabaña et al. Ann. ZOOL. Fennici Vol. 51 A C F B A P R Q D M L K B N O 46 H G I J E Fig. 3. Networks based on the matrix of (A) mtdna (ND4) and (B) ndna (ACA4). White circles and slices represent specimens identified as T. merianae; black circles and slices represent specimens identified as T. rufescens. Each haplotype or allele is indicated by a circle and coded with the same letter as in Table 1. The size of each circle represents the frequency of the denoted haplotype or allele. The dots in the branches that connect the circles represent numbers of mutations separating the different haplotypes and alleles. The five individuals with introgressed haplotypes presented in both networks the most frequent variants (Table 1 and Fig. 3). None of the specimens presented ndna alleles belonging to both parental species, indicating that F 1 hybrids were not found in this study. Discussion In this study, we present the first evidence of natural hybridization between species of Tupinambis: some individuals were grouped in different clades, depending on whether the phylogenetic trees and/or haplotype network were built based on the mitochondrial or nuclear dataset. The inconsistency in phylogenetic estimates from different sources of evidence (mtdna and ndna) has been interpreted in many taxa as the result of hybridization (Seehausen 2004). Another possible explanation could be an incomplete lineage sorting, which means that some individuals of these two species present identical or very similar haplotypes due to common ancestry and the short time elapsed since the separation between them. However, the identified hybrids occur only in the sympatry zone and are not randomly distributed across the entire study area, as expected if they were the result of common ancestry. Therefore, current gene flow between T. merianae and T. rufescens in the sympatric zone is the most likely explanation. As mentioned above, Cei (1993) suggested the existence of a hybrid individual between the two studied species based on its scalation pattern. Fitzgerald et al. (1999) also proposed the possibility of introgression of mitochondrial DNA between T. rufescens and T duseni to explain the incongruence between molecular and morphological data. However, in their study the specimens of T. duseni that presented mtdna of T. rufescens did not occur in sympatry with the latter species; for this reason, the authors considered introgression as a less likely explanation for their results. In our study, the use of both mtdna and ndna markers analyzing specimens of T. merianae and T. rufecens in areas of sympatry and allopatry provides the first evidence of the existence of hybridization in the genus. The presence of the mitochondrial gene sequences of T. rufescens in four T. merianae individuals suggests that females of the former
8 Ann. Zool. Fennici Vol. 51 Hybridization in Tupinambis 347 species mate with males of the latter. On the other hand, the fact that a specimen of T. rufescens presented a T. merianae mitochondrial haplotype indicates that hybridization can also occur in the opposite direction; thus, the process would be reciprocal. Hybrid individuals detected originated from backcrossing (no F 1 hybrids were found), demonstrating the occurrence of introgression between T. merianae and T. rufecens. Furthermore, mitochondrial haplotypes found in the hybrid specimens were different, suggesting multiple hybridization events. Bolnick and Near (2005), in a study of fishes of the Centrachidae family established that natural hybridization is common even among taxa that have been separated for up to 14 mya. Péres (2003) estimated that the divergence between T. merianae and T. rufescens occurred during the late Miocene (10 mya). According to our results, this amount of time would not have been enough to reach a degree of reproductive isolation that prevents backcrossing between these species. The hybrid zone corresponding to the area of sympatry between T. merianae and T. rufescens is the ecotone between Arid Chaco and Espinal (Cardozo et al. 2012). However, due to the low number of hybrids detected we cannot establish other characteristics of this zone, such as size, or if it corresponds to a continuous region across the ecotone or to a mosaic of patches. The finding of natural hybridization in populations of Tupinambis spp. raises many questions about how reproductive strategies, patterns of dispersal and the ecological characteristics of hybrids shape the hybrid zone influencing the evolution of the lineages. New studies using highly variable molecular markers, microsatellites, might answer these questions. Acknowledgments We are grateful to the local people for their invaluable assistance in data collection. We also thank the members of Laboratorio de Biología del Comportamiento, Sergio Naretto, Cecilia Blengini, Gabriela Cardozo and Valeria Di Cola, for field assistance and comments that helped to improve this manuscript. We also thank two anonymous reviewers for their careful reading of the first draft of the manuscript and their constructive comments and suggestions. Our work was funded by Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET), Fondo para la Investigación Científica y Tecnológica (FONCyT), MinCyT Córdoba-Préstamo BID-PID no. 013/2009, Secretaría de Ciencia y Tecnología (SeCyT) and Universidad Nacional de Córdoba, Argentina. References Anderson, E. 1949: Introgressive hybridization. J. Wiley, New York. Bandelt, H. J., Forster, P. & Röhl, A. 1999: Median-joining networks for inferring intraspecific phylogenies. Molecular Biology and Evolution 16: Barton, N. H. 2001: The role of hybridization in evolution. Molecular Ecology 10: Bolnick, D. I. & Near, T. J. 2005: Tempo of hybrid inviability in Centrarchid fishes (Teleostei: Centrarchidae). Evolution 59: Bruford, M. E., Hanotte, O., Brookfield, J. F. Y. & Burke, T. 1992: Single-locus and multilocus DNA fingerprinting. In: Hoelzel, A. R. (ed.), Molecular genetic analysis of populations: a practical approach: Oxford University Press, New York. Cardozo, G., Naretto, S., Zak, M. & Chiaraviglio, M. 2012: The Role of Landscape in Contact Zones of Sister Species of Lizard. In: Tiefenbacher, J. (ed.), Perspectives on nature conservation patterns, pressures and prospects: Ed. Intech, Croatia. Cei, J. M. 1986: Reptiles del centro, centro-oeste y sur de Argentina. Herpetofauna de las Zonas Áridas y Semiáridas. Monografia VI, Museo Regionale de Scienze Naturali, Torino. Cei, J. M. 1993: Reptiles del noroeste, nordeste y este de la Argentina, Herpetofauna de las selvas subtropicales, Puna y Pampas. Monografia XIV, Museo Regionale di Scienze Naturali, Torino. Cole, C. J., Painter, C. W., Dessauer, H. C. & Taylor, H. L. 2007: Hybridization between the endangered unisexual gray-checkered whiptail lizard (Aspidoscelis dixoni) and the bisexual western whiptail lizard (Aspidoscelis tigris) in southwestern New Mexico. American Museum Novitates 3555: Dessauer, H. C., Cole, C. J. & Townsend, C. R. 2000: Hybridization among western whiptail lizards (Cnemidophorus tigris) in southwestern New Mexico: population genetics, morphology, and ecology in three contact zones. Bulletin of the American Museum of Natural History 246: Dowling T. E. & Secor, C. L. 1997: The role of Hybridization and introgression in the diversification in animals. Annual Review of Ecology and Systematic 28: Edgar, R. C. 2004: MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Research 32: Fitzgerald, L. A., Cook, J. A. & Aquino, A. L. 1999: Molecular phylogenetics and conservation of Tupinambis (Sauria: Teiidae). Copeia 1999: Forstner, M., Davis, S. & Arévalo, E. 1995: Support for the
9 348 Cabaña et al. Ann. ZOOL. Fennici Vol. 51 hypothesis of anguimorph ancestry for the suborder serpentes from phylogenetic analysis of mitochondrial DNA sequences. Molecular Phylogenetics and Evolution 4: Giffor, M. E. & Larson, A. 2008: In situ genetic differentiation in a Hispaniolan lizard (Ameiva chrysolaema): A multilocus perspective. Molecular Phylogenetics and Evolution 49: Giugliano, L., Collevatti, R. & Colli, G. R. 2007: Molecular dating and phylogenetic relationships among Teiidae (Squamata) inferred by molecular and morphological data. Molecular Phylogenetics and Evolution 45: Leache, A. & McGuire, J. 2006: Phylogenetic relationships of horned lizards (Phrynosoma) based on nuclear and mitochondrial data: evidence for a misleading mitochondrial gene tree. Molecular Phylogenetics and Evolution 39: Mallet, J. 2007: Hybrid speciation. Nature 446: Mallet, J. 2008: Hybridization, ecological races and the nature of species: empirical evidence for the ease of speciation. Philosophical Transactions of the Royal Society B 363: Manríquez-Morán, N. L. 2007: Diversidad clonal en los lacertilios unisexuales del género Aspidoscelis. Boletín de la Sociedad Herpetológica Mexicana 15: Martínez, J. J., González-Ittig, R.E., Theiler, G. R., Ojeda, R., Lanzone, C., Ojeda, A. & Gardenal, C. N. 2009: Patterns of speciation in two sibling species of Graomys (Rodentia, Cricetidae) based on mtdna sequences. Journal of Zoological Systematics and Evolutionary Research 48: Péres, A. K. Jr. 2003: Teiid lizards of the genus Tupinambis: taxonomic notes, geographic review, and a key to the species. Ph.D. thesis, Universidade de Brasília. Pinho, C., Harris, D. J. & Ferrand, N. 2007: Contrasting patterns of population subdivision and historical demography in three western Mediterranean lizard species inferred from mitochondrial DNA variation. Molecular Ecology 6: Porini, G. 2006: Proyecto Tupinambis. Una propuesta para el manejo de Tupinambis rufescens y Tupinambis merianae en la Argentina. In: Bolkovic, M. & Ramadori, D. (eds.), Manejo de Fauna Silvestre en la Argentina: Programa de Uso Sustentable, Dirección de Fauna Silvestre, Secretaría de Ambiente y Desarrollo Sustentable, Buenos Aires. Posada, D. 2008: jmodeltest: phylogenetic model averaging. Molecular Biology and Evolution 25: Reeder, T. W., Cole, C. J. & Dessauer, H. C. 2002: Phylogenetic relationships of whiptail lizards of the genus Cnemidophorus (Squamata: Teiidae): a test of monophyly, reevaluation of karyotypic evolution, and review of hybrid origins. American Museum Novitates 3365: Ronquist, F. & Huelsenbeck, J. P. 2003: MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19: Seehausen, O. 2004: Hybridization and adaptative radiation. Trends in Ecology and Evolution 19: Stephens, M. & Donnelly, P. 2003: A comparison of Bayesian methods for haplotype reconstruction from population genotype data. The American Journal of Human Genetics 73: Swofford, D. L. 1998: PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. Sunderland, MA: Sinauer Associates. Taylor, H. L., Cole, C. J., Hardy, L. M., Dessauer, H. C., Townsend, C. R., Walker, J. M. & Cordes, J. E. 2001: Natural hybridization between the teiid lizards Cnemidophorus tesselatus (parthenogenetic) and C. tigris marmoratus (bisexual): assessment of evolutionary alternatives. American Museum Novitates 3345: Taylor, H. L., Lemos-espinal, J. A. & Smith, H. M. 2003: Morphological characteristics of a newly discovered population of Aspidoscelis tesselata (Squamata: Teiidae) from Chihuahua, México, the identity of an associated hybrid, and a pattern of geographic variation. The Southwestern Naturalist 48: Waltari, E. & Edwards, S. V. 2002: Evolutionary dynamics of intron size, genome size, and physiological correlated in Archosaurs. The American Naturalist 160: Wirtz, P. 1999: Mother species father species: unidirectional hybridization in animals with female choice. Animal Behavior 58: Zarza, E., Reynoso, V. & Emerson, E. 2008: Diversification in the northern neotropics: mitochondrial and nuclear DNA phylogeography of the iguana Ctenosaura pectinata and related species. Molecular Ecology 17: This article is also available at
Lecture 11 Wednesday, September 19, 2012
 Lecture 11 Wednesday, September 19, 2012 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean
Lecture 11 Wednesday, September 19, 2012 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean
CLADISTICS Student Packet SUMMARY Phylogeny Phylogenetic trees/cladograms
 CLADISTICS Student Packet SUMMARY PHYLOGENETIC TREES AND CLADOGRAMS ARE MODELS OF EVOLUTIONARY HISTORY THAT CAN BE TESTED Phylogeny is the history of descent of organisms from their common ancestor. Phylogenetic
CLADISTICS Student Packet SUMMARY PHYLOGENETIC TREES AND CLADOGRAMS ARE MODELS OF EVOLUTIONARY HISTORY THAT CAN BE TESTED Phylogeny is the history of descent of organisms from their common ancestor. Phylogenetic
GEODIS 2.0 DOCUMENTATION
 GEODIS.0 DOCUMENTATION 1999-000 David Posada and Alan Templeton Contact: David Posada, Department of Zoology, 574 WIDB, Provo, UT 8460-555, USA Fax: (801) 78 74 e-mail: dp47@email.byu.edu 1. INTRODUCTION
GEODIS.0 DOCUMENTATION 1999-000 David Posada and Alan Templeton Contact: David Posada, Department of Zoology, 574 WIDB, Provo, UT 8460-555, USA Fax: (801) 78 74 e-mail: dp47@email.byu.edu 1. INTRODUCTION
PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, NY 10024
 PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, NY 10024 Number 3365, 61 pp., 7 figures, 3 tables May 17, 2002 Phylogenetic Relationships of Whiptail Lizards
PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, NY 10024 Number 3365, 61 pp., 7 figures, 3 tables May 17, 2002 Phylogenetic Relationships of Whiptail Lizards
INQUIRY & INVESTIGATION
 INQUIRY & INVESTIGTION Phylogenies & Tree-Thinking D VID. UM SUSN OFFNER character a trait or feature that varies among a set of taxa (e.g., hair color) character-state a variant of a character that occurs
INQUIRY & INVESTIGTION Phylogenies & Tree-Thinking D VID. UM SUSN OFFNER character a trait or feature that varies among a set of taxa (e.g., hair color) character-state a variant of a character that occurs
6. The lifetime Darwinian fitness of one organism is greater than that of another organism if: A. it lives longer than the other B. it is able to outc
 1. The money in the kingdom of Florin consists of bills with the value written on the front, and pictures of members of the royal family on the back. To test the hypothesis that all of the Florinese $5
1. The money in the kingdom of Florin consists of bills with the value written on the front, and pictures of members of the royal family on the back. To test the hypothesis that all of the Florinese $5
History of Lineages. Chapter 11. Jamie Oaks 1. April 11, Kincaid Hall 524. c 2007 Boris Kulikov boris-kulikov.blogspot.
 History of Lineages Chapter 11 Jamie Oaks 1 1 Kincaid Hall 524 joaks1@gmail.com April 11, 2014 c 2007 Boris Kulikov boris-kulikov.blogspot.com History of Lineages J. Oaks, University of Washington 1/46
History of Lineages Chapter 11 Jamie Oaks 1 1 Kincaid Hall 524 joaks1@gmail.com April 11, 2014 c 2007 Boris Kulikov boris-kulikov.blogspot.com History of Lineages J. Oaks, University of Washington 1/46
Rediscovered population of Mexican Plateau spotted whiptail lizard, Aspidoscelis septemvittata (Teiidae), from México, D.F.
 Western North American Naturalist Volume 69 Number 1 Article 6 4-24-2009 Rediscovered population of Mexican Plateau spotted whiptail lizard, Aspidoscelis septemvittata (Teiidae), from México, D.F. Oswaldo
Western North American Naturalist Volume 69 Number 1 Article 6 4-24-2009 Rediscovered population of Mexican Plateau spotted whiptail lizard, Aspidoscelis septemvittata (Teiidae), from México, D.F. Oswaldo
Species: Panthera pardus Genus: Panthera Family: Felidae Order: Carnivora Class: Mammalia Phylum: Chordata
 CHAPTER 6: PHYLOGENY AND THE TREE OF LIFE AP Biology 3 PHYLOGENY AND SYSTEMATICS Phylogeny - evolutionary history of a species or group of related species Systematics - analytical approach to understanding
CHAPTER 6: PHYLOGENY AND THE TREE OF LIFE AP Biology 3 PHYLOGENY AND SYSTEMATICS Phylogeny - evolutionary history of a species or group of related species Systematics - analytical approach to understanding
UNIT III A. Descent with Modification(Ch19) B. Phylogeny (Ch20) C. Evolution of Populations (Ch21) D. Origin of Species or Speciation (Ch22)
 UNIT III A. Descent with Modification(Ch9) B. Phylogeny (Ch2) C. Evolution of Populations (Ch2) D. Origin of Species or Speciation (Ch22) Classification in broad term simply means putting things in classes
UNIT III A. Descent with Modification(Ch9) B. Phylogeny (Ch2) C. Evolution of Populations (Ch2) D. Origin of Species or Speciation (Ch22) Classification in broad term simply means putting things in classes
A Mitochondrial DNA Phylogeny of Extant Species of the Genus Trachemys with Resulting Taxonomic Implications
 NOTES AND FIELD REPORTS 131 Chelonian Conservation and Biology, 2008, 7(1): 131 135 Ó 2008 Chelonian Research Foundation A Mitochondrial DNA Phylogeny of Extant Species of the Genus Trachemys with Resulting
NOTES AND FIELD REPORTS 131 Chelonian Conservation and Biology, 2008, 7(1): 131 135 Ó 2008 Chelonian Research Foundation A Mitochondrial DNA Phylogeny of Extant Species of the Genus Trachemys with Resulting
Modern Evolutionary Classification. Lesson Overview. Lesson Overview Modern Evolutionary Classification
 Lesson Overview 18.2 Modern Evolutionary Classification THINK ABOUT IT Darwin s ideas about a tree of life suggested a new way to classify organisms not just based on similarities and differences, but
Lesson Overview 18.2 Modern Evolutionary Classification THINK ABOUT IT Darwin s ideas about a tree of life suggested a new way to classify organisms not just based on similarities and differences, but
PARTIAL REPORT. Juvenile hybrid turtles along the Brazilian coast RIO GRANDE FEDERAL UNIVERSITY
 RIO GRANDE FEDERAL UNIVERSITY OCEANOGRAPHY INSTITUTE MARINE MOLECULAR ECOLOGY LABORATORY PARTIAL REPORT Juvenile hybrid turtles along the Brazilian coast PROJECT LEADER: MAIRA PROIETTI PROFESSOR, OCEANOGRAPHY
RIO GRANDE FEDERAL UNIVERSITY OCEANOGRAPHY INSTITUTE MARINE MOLECULAR ECOLOGY LABORATORY PARTIAL REPORT Juvenile hybrid turtles along the Brazilian coast PROJECT LEADER: MAIRA PROIETTI PROFESSOR, OCEANOGRAPHY
The Making of the Fittest: LESSON STUDENT MATERIALS USING DNA TO EXPLORE LIZARD PHYLOGENY
 The Making of the Fittest: Natural The The Making Origin Selection of the of Species and Fittest: Adaptation Natural Lizards Selection in an Evolutionary and Adaptation Tree INTRODUCTION USING DNA TO EXPLORE
The Making of the Fittest: Natural The The Making Origin Selection of the of Species and Fittest: Adaptation Natural Lizards Selection in an Evolutionary and Adaptation Tree INTRODUCTION USING DNA TO EXPLORE
Bi156 Lecture 1/13/12. Dog Genetics
 Bi156 Lecture 1/13/12 Dog Genetics The radiation of the family Canidae occurred about 100 million years ago. Dogs are most closely related to wolves, from which they diverged through domestication about
Bi156 Lecture 1/13/12 Dog Genetics The radiation of the family Canidae occurred about 100 million years ago. Dogs are most closely related to wolves, from which they diverged through domestication about
Testing Phylogenetic Hypotheses with Molecular Data 1
 Testing Phylogenetic Hypotheses with Molecular Data 1 How does an evolutionary biologist quantify the timing and pathways for diversification (speciation)? If we observe diversification today, the processes
Testing Phylogenetic Hypotheses with Molecular Data 1 How does an evolutionary biologist quantify the timing and pathways for diversification (speciation)? If we observe diversification today, the processes
Inferring Ancestor-Descendant Relationships in the Fossil Record
 Inferring Ancestor-Descendant Relationships in the Fossil Record (With Statistics) David Bapst, Melanie Hopkins, April Wright, Nick Matzke & Graeme Lloyd GSA 2016 T151 Wednesday Sept 28 th, 9:15 AM Feel
Inferring Ancestor-Descendant Relationships in the Fossil Record (With Statistics) David Bapst, Melanie Hopkins, April Wright, Nick Matzke & Graeme Lloyd GSA 2016 T151 Wednesday Sept 28 th, 9:15 AM Feel
Title: Phylogenetic Methods and Vertebrate Phylogeny
 Title: Phylogenetic Methods and Vertebrate Phylogeny Central Question: How can evolutionary relationships be determined objectively? Sub-questions: 1. What affect does the selection of the outgroup have
Title: Phylogenetic Methods and Vertebrate Phylogeny Central Question: How can evolutionary relationships be determined objectively? Sub-questions: 1. What affect does the selection of the outgroup have
Introduction to phylogenetic trees and tree-thinking Copyright 2005, D. A. Baum (Free use for non-commercial educational pruposes)
 Introduction to phylogenetic trees and tree-thinking Copyright 2005, D. A. Baum (Free use for non-commercial educational pruposes) Phylogenetics is the study of the relationships of organisms to each other.
Introduction to phylogenetic trees and tree-thinking Copyright 2005, D. A. Baum (Free use for non-commercial educational pruposes) Phylogenetics is the study of the relationships of organisms to each other.
Genetic homogeneity between two populations of the parthenogenetic lizard Aspidoscelis cozumela
 Revista Mexicana de Biodiversidad 79: 421-426, 2008 Genetic homogeneity between two populations of the parthenogenetic lizard Aspidoscelis cozumela Homogeneidad genética entre dos poblaciones de la lagartija
Revista Mexicana de Biodiversidad 79: 421-426, 2008 Genetic homogeneity between two populations of the parthenogenetic lizard Aspidoscelis cozumela Homogeneidad genética entre dos poblaciones de la lagartija
Phylogeographic assessment of Acanthodactylus boskianus (Reptilia: Lacertidae) based on phylogenetic analysis of mitochondrial DNA.
 Zoology Department Phylogeographic assessment of Acanthodactylus boskianus (Reptilia: Lacertidae) based on phylogenetic analysis of mitochondrial DNA By HAGAR IBRAHIM HOSNI BAYOUMI A thesis submitted in
Zoology Department Phylogeographic assessment of Acanthodactylus boskianus (Reptilia: Lacertidae) based on phylogenetic analysis of mitochondrial DNA By HAGAR IBRAHIM HOSNI BAYOUMI A thesis submitted in
A new karyotypic formula for the genus Amphisbaena (Squamata: Amphisbaenidae)
 Phyllomedusa 9(1):75-80, 2010 2010 Departamento de Ciências Biológicas - ESALQ - USP ISSN 1519-1397 Short Communication A new karyotypic formula for the genus Amphisbaena (Squamata: Amphisbaenidae) Camila
Phyllomedusa 9(1):75-80, 2010 2010 Departamento de Ciências Biológicas - ESALQ - USP ISSN 1519-1397 Short Communication A new karyotypic formula for the genus Amphisbaena (Squamata: Amphisbaenidae) Camila
Prof. Neil. J.L. Heideman
 Prof. Neil. J.L. Heideman Position Office Mailing address E-mail : Vice-dean (Professor of Zoology) : No. 10, Biology Building : P.O. Box 339 (Internal Box 44), Bloemfontein 9300, South Africa : heidemannj.sci@mail.uovs.ac.za
Prof. Neil. J.L. Heideman Position Office Mailing address E-mail : Vice-dean (Professor of Zoology) : No. 10, Biology Building : P.O. Box 339 (Internal Box 44), Bloemfontein 9300, South Africa : heidemannj.sci@mail.uovs.ac.za
Bio 1B Lecture Outline (please print and bring along) Fall, 2006
 Bio 1B Lecture Outline (please print and bring along) Fall, 2006 B.D. Mishler, Dept. of Integrative Biology 2-6810, bmishler@berkeley.edu Evolution lecture #4 -- Phylogenetic Analysis (Cladistics) -- Oct.
Bio 1B Lecture Outline (please print and bring along) Fall, 2006 B.D. Mishler, Dept. of Integrative Biology 2-6810, bmishler@berkeley.edu Evolution lecture #4 -- Phylogenetic Analysis (Cladistics) -- Oct.
Natural hybridization of the bisexual teiid lizard Cnemidophorus inornatus and the unisexual Cnemidophorus perplexus in southern New Mexico
 University of Colorado, Boulder CU Scholar Series in Biology Ecology & Evolutionary Biology Winter 3-1-1966 Natural hybridization of the bisexual teiid lizard Cnemidophorus inornatus and the unisexual
University of Colorado, Boulder CU Scholar Series in Biology Ecology & Evolutionary Biology Winter 3-1-1966 Natural hybridization of the bisexual teiid lizard Cnemidophorus inornatus and the unisexual
17.2 Classification Based on Evolutionary Relationships Organization of all that speciation!
 Organization of all that speciation! Patterns of evolution.. Taxonomy gets an over haul! Using more than morphology! 3 domains, 6 kingdoms KEY CONCEPT Modern classification is based on evolutionary relationships.
Organization of all that speciation! Patterns of evolution.. Taxonomy gets an over haul! Using more than morphology! 3 domains, 6 kingdoms KEY CONCEPT Modern classification is based on evolutionary relationships.
Partial island submergence and speciation in an adaptive radiation: a multilocus analysis of the Cuban green anoles
 Received 2 May 2004 Accepted 27 May 2004 Published online 25 October 2004 Partial island submergence and speciation in an adaptive radiation: a multilocus analysis of the Cuban green anoles Richard E.
Received 2 May 2004 Accepted 27 May 2004 Published online 25 October 2004 Partial island submergence and speciation in an adaptive radiation: a multilocus analysis of the Cuban green anoles Richard E.
The Rufford Foundation Final Report
 The Rufford Foundation Final Report Congratulations on the completion of your project that was supported by The Rufford Foundation. We ask all grant recipients to complete a Final Report Form that helps
The Rufford Foundation Final Report Congratulations on the completion of your project that was supported by The Rufford Foundation. We ask all grant recipients to complete a Final Report Form that helps
Range extension of the critically endangered true poison-dart frog, Phyllobates terribilis (Anura: Dendrobatidae), in western Colombia
 Acta Herpetologica 7(2): 365-x, 2012 Range extension of the critically endangered true poison-dart frog, Phyllobates terribilis (Anura: Dendrobatidae), in western Colombia Roberto Márquez 1, *, Germán
Acta Herpetologica 7(2): 365-x, 2012 Range extension of the critically endangered true poison-dart frog, Phyllobates terribilis (Anura: Dendrobatidae), in western Colombia Roberto Márquez 1, *, Germán
Comparing DNA Sequences Cladogram Practice
 Name Period Assignment # See lecture questions 75, 122-123, 127, 137 Comparing DNA Sequences Cladogram Practice BACKGROUND Between 1990 2003, scientists working on an international research project known
Name Period Assignment # See lecture questions 75, 122-123, 127, 137 Comparing DNA Sequences Cladogram Practice BACKGROUND Between 1990 2003, scientists working on an international research project known
2013 Holiday Lectures on Science Medicine in the Genomic Era
 INTRODUCTION Figure 1. Tasha. Scientists sequenced the first canine genome using DNA from a boxer named Tasha. Meet Tasha, a boxer dog (Figure 1). In 2005, scientists obtained the first complete dog genome
INTRODUCTION Figure 1. Tasha. Scientists sequenced the first canine genome using DNA from a boxer named Tasha. Meet Tasha, a boxer dog (Figure 1). In 2005, scientists obtained the first complete dog genome
Phylogeny Reconstruction
 Phylogeny Reconstruction Trees, Methods and Characters Reading: Gregory, 2008. Understanding Evolutionary Trees (Polly, 2006) Lab tomorrow Meet in Geology GY522 Bring computers if you have them (they will
Phylogeny Reconstruction Trees, Methods and Characters Reading: Gregory, 2008. Understanding Evolutionary Trees (Polly, 2006) Lab tomorrow Meet in Geology GY522 Bring computers if you have them (they will
Phylogeography and diversification history of the day-gecko genus Phelsuma in the Seychelles islands. Rocha et al.
 Phylogeography and diversification history of the day-gecko genus Phelsuma in the Seychelles islands Rocha et al. Rocha et al. BMC Evolutionary Biology 2013, 13:3 Rocha et al. BMC Evolutionary Biology
Phylogeography and diversification history of the day-gecko genus Phelsuma in the Seychelles islands Rocha et al. Rocha et al. BMC Evolutionary Biology 2013, 13:3 Rocha et al. BMC Evolutionary Biology
Temporal mitochondrial DNA variation in honeybee populations from Tenerife (Canary Islands, Spain)
 Temporal mitochondrial DNA variation in honeybee populations from Tenerife (Canary Islands, Spain) Mª Jesús Madrid-Jiménez, Irene Muñoz, Pilar De la Rúa Dpto. de Zoología y Antropología Física, Facultad
Temporal mitochondrial DNA variation in honeybee populations from Tenerife (Canary Islands, Spain) Mª Jesús Madrid-Jiménez, Irene Muñoz, Pilar De la Rúa Dpto. de Zoología y Antropología Física, Facultad
PUBLICATIONS (PEER REVIEWED)
 Matthew E. Gifford EDUCATION Present Washington University, Department of Biology Campus Box 1137, St. Louis, Missouri 63130 Office: (314)935 5302, Cell: (314)550 0485, Email: gifford@biology2.wustl.edu
Matthew E. Gifford EDUCATION Present Washington University, Department of Biology Campus Box 1137, St. Louis, Missouri 63130 Office: (314)935 5302, Cell: (314)550 0485, Email: gifford@biology2.wustl.edu
Cladistics (reading and making of cladograms)
 Cladistics (reading and making of cladograms) Definitions Systematics The branch of biological sciences concerned with classifying organisms Taxon (pl: taxa) Any unit of biological diversity (eg. Animalia,
Cladistics (reading and making of cladograms) Definitions Systematics The branch of biological sciences concerned with classifying organisms Taxon (pl: taxa) Any unit of biological diversity (eg. Animalia,
Darwin s Finches: A Thirty Year Study.
 Darwin s Finches: A Thirty Year Study. I. Mit-DNA Based Phylogeny (Figure 1). 1. All Darwin s finches descended from South American grassquit (small finch) ancestor circa 3 Mya. 2. Galapagos colonized
Darwin s Finches: A Thirty Year Study. I. Mit-DNA Based Phylogeny (Figure 1). 1. All Darwin s finches descended from South American grassquit (small finch) ancestor circa 3 Mya. 2. Galapagos colonized
1 This question is about the evolution, genetics, behaviour and physiology of cats.
 1 This question is about the evolution, genetics, behaviour and physiology of cats. Fig. 1.1 (on the insert) shows a Scottish wildcat, Felis sylvestris. Modern domestic cats evolved from a wild ancestor
1 This question is about the evolution, genetics, behaviour and physiology of cats. Fig. 1.1 (on the insert) shows a Scottish wildcat, Felis sylvestris. Modern domestic cats evolved from a wild ancestor
National Finch & Softbill Society
 First Class Mail U.S. Postage PAID Shawnee Msn KS Permit No. 84! 21 Oakcrest Rd S. Weymouth, MA 02190 Journal of the National Finch & Softbill Society Vol. 28, No. 4 Jul / Aug 2011 Using Genetics to Understand
First Class Mail U.S. Postage PAID Shawnee Msn KS Permit No. 84! 21 Oakcrest Rd S. Weymouth, MA 02190 Journal of the National Finch & Softbill Society Vol. 28, No. 4 Jul / Aug 2011 Using Genetics to Understand
Ch 1.2 Determining How Species Are Related.notebook February 06, 2018
 Name 3 "Big Ideas" from our last notebook lecture: * * * 1 WDYR? Of the following organisms, which is the closest relative of the "Snowy Owl" (Bubo scandiacus)? a) barn owl (Tyto alba) b) saw whet owl
Name 3 "Big Ideas" from our last notebook lecture: * * * 1 WDYR? Of the following organisms, which is the closest relative of the "Snowy Owl" (Bubo scandiacus)? a) barn owl (Tyto alba) b) saw whet owl
Phylogenetic relationships of horned lizards (Phrynosoma) based on nuclear and mitochondrial data: Evidence for a misleading mitochondrial gene tree
 Molecular Phylogenetics and Evolution 39 (2006) 628 644 www.elsevier.com/locate/ympev Phylogenetic relationships of horned lizards (Phrynosoma) based on nuclear and mitochondrial data: Evidence for a misleading
Molecular Phylogenetics and Evolution 39 (2006) 628 644 www.elsevier.com/locate/ympev Phylogenetic relationships of horned lizards (Phrynosoma) based on nuclear and mitochondrial data: Evidence for a misleading
Systematics and taxonomy of the genus Culicoides what is coming next?
 Systematics and taxonomy of the genus Culicoides what is coming next? Claire Garros 1, Bruno Mathieu 2, Thomas Balenghien 1, Jean-Claude Delécolle 2 1 CIRAD, Montpellier, France 2 IPPTS, Strasbourg, France
Systematics and taxonomy of the genus Culicoides what is coming next? Claire Garros 1, Bruno Mathieu 2, Thomas Balenghien 1, Jean-Claude Delécolle 2 1 CIRAD, Montpellier, France 2 IPPTS, Strasbourg, France
LABORATORY EXERCISE 7: CLADISTICS I
 Biology 4415/5415 Evolution LABORATORY EXERCISE 7: CLADISTICS I Take a group of organisms. Let s use five: a lungfish, a frog, a crocodile, a flamingo, and a human. How to reconstruct their relationships?
Biology 4415/5415 Evolution LABORATORY EXERCISE 7: CLADISTICS I Take a group of organisms. Let s use five: a lungfish, a frog, a crocodile, a flamingo, and a human. How to reconstruct their relationships?
Multi-Locus Phylogeographic and Population Genetic Analysis of Anolis carolinensis: Historical Demography of a Genomic Model Species
 City University of New York (CUNY) CUNY Academic Works Publications and Research Queens College June 2012 Multi-Locus Phylogeographic and Population Genetic Analysis of Anolis carolinensis: Historical
City University of New York (CUNY) CUNY Academic Works Publications and Research Queens College June 2012 Multi-Locus Phylogeographic and Population Genetic Analysis of Anolis carolinensis: Historical
TOPIC CLADISTICS
 TOPIC 5.4 - CLADISTICS 5.4 A Clades & Cladograms https://upload.wikimedia.org/wikipedia/commons/thumb/4/46/clade-grade_ii.svg IB BIO 5.4 3 U1: A clade is a group of organisms that have evolved from a common
TOPIC 5.4 - CLADISTICS 5.4 A Clades & Cladograms https://upload.wikimedia.org/wikipedia/commons/thumb/4/46/clade-grade_ii.svg IB BIO 5.4 3 U1: A clade is a group of organisms that have evolved from a common
COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST
 Big Idea 1 Evolution INVESTIGATION 3 COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST How can bioinformatics be used as a tool to determine evolutionary relationships and to
Big Idea 1 Evolution INVESTIGATION 3 COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST How can bioinformatics be used as a tool to determine evolutionary relationships and to
Global comparisons of beta diversity among mammals, birds, reptiles, and amphibians across spatial scales and taxonomic ranks
 Journal of Systematics and Evolution 47 (5): 509 514 (2009) doi: 10.1111/j.1759-6831.2009.00043.x Global comparisons of beta diversity among mammals, birds, reptiles, and amphibians across spatial scales
Journal of Systematics and Evolution 47 (5): 509 514 (2009) doi: 10.1111/j.1759-6831.2009.00043.x Global comparisons of beta diversity among mammals, birds, reptiles, and amphibians across spatial scales
A range-wide synthesis and timeline for phylogeographic events in the red fox (Vulpes vulpes)
 Kutschera et al. BMC Evolutionary Biology 2013, 13:114 RESEARCH ARTICLE Open Access A range-wide synthesis and timeline for phylogeographic events in the red fox (Vulpes vulpes) Verena E Kutschera 1*,
Kutschera et al. BMC Evolutionary Biology 2013, 13:114 RESEARCH ARTICLE Open Access A range-wide synthesis and timeline for phylogeographic events in the red fox (Vulpes vulpes) Verena E Kutschera 1*,
BioSci 110, Fall 08 Exam 2
 1. is the cell division process that results in the production of a. mitosis; 2 gametes b. meiosis; 2 gametes c. meiosis; 2 somatic (body) cells d. mitosis; 4 somatic (body) cells e. *meiosis; 4 gametes
1. is the cell division process that results in the production of a. mitosis; 2 gametes b. meiosis; 2 gametes c. meiosis; 2 somatic (body) cells d. mitosis; 4 somatic (body) cells e. *meiosis; 4 gametes
The impact of the recognizing evolution on systematics
 The impact of the recognizing evolution on systematics 1. Genealogical relationships between species could serve as the basis for taxonomy 2. Two sources of similarity: (a) similarity from descent (b)
The impact of the recognizing evolution on systematics 1. Genealogical relationships between species could serve as the basis for taxonomy 2. Two sources of similarity: (a) similarity from descent (b)
Fig Phylogeny & Systematics
 Fig. 26- Phylogeny & Systematics Tree of Life phylogenetic relationship for 3 clades (http://evolution.berkeley.edu Fig. 26-2 Phylogenetic tree Figure 26.3 Taxonomy Taxon Carolus Linnaeus Species: Panthera
Fig. 26- Phylogeny & Systematics Tree of Life phylogenetic relationship for 3 clades (http://evolution.berkeley.edu Fig. 26-2 Phylogenetic tree Figure 26.3 Taxonomy Taxon Carolus Linnaeus Species: Panthera
Volume 2 Number 1, July 2012 ISSN:
 Volume 2 Number 1, July 2012 ISSN: 229-9769 Published by Faculty of Resource Science and Technology Borneo J. Resour. Sci. Tech. (2012) 2: 20-27 Molecular Phylogeny of Sarawak Green Sea Turtle (Chelonia
Volume 2 Number 1, July 2012 ISSN: 229-9769 Published by Faculty of Resource Science and Technology Borneo J. Resour. Sci. Tech. (2012) 2: 20-27 Molecular Phylogeny of Sarawak Green Sea Turtle (Chelonia
USING DNA TO EXPLORE LIZARD PHYLOGENY
 Species The MThe aking of the offittest: The Making of the Fittest: in anand Natural Selection Adaptation Tree Natural Selection and Adaptation USING DNA TO EXPLORE LIZARD PHYLOGENY OVERVIEW This lesson
Species The MThe aking of the offittest: The Making of the Fittest: in anand Natural Selection Adaptation Tree Natural Selection and Adaptation USING DNA TO EXPLORE LIZARD PHYLOGENY OVERVIEW This lesson
Old age, multiple formations or genetic plasticity? Clonal diversity in the uniparental Caucasian rock lizard, Lacerta dahli
 Genetica 101: 125 130, 1997. 125 c 1997 Kluwer Academic Publishers. Printed in the Netherlands. Old age, multiple formations or genetic plasticity? Clonal diversity in the uniparental Caucasian rock lizard,
Genetica 101: 125 130, 1997. 125 c 1997 Kluwer Academic Publishers. Printed in the Netherlands. Old age, multiple formations or genetic plasticity? Clonal diversity in the uniparental Caucasian rock lizard,
RESEARCH REPOSITORY.
 RESEARCH REPOSITORY This is the author s final version of the work, as accepted for publication following peer review but without the publisher s layout or pagination. The definitive version is available
RESEARCH REPOSITORY This is the author s final version of the work, as accepted for publication following peer review but without the publisher s layout or pagination. The definitive version is available
Reintroducing bettongs to the ACT: issues relating to genetic diversity and population dynamics The guest speaker at NPA s November meeting was April
 Reintroducing bettongs to the ACT: issues relating to genetic diversity and population dynamics The guest speaker at NPA s November meeting was April Suen, holder of NPA s 2015 scholarship for honours
Reintroducing bettongs to the ACT: issues relating to genetic diversity and population dynamics The guest speaker at NPA s November meeting was April Suen, holder of NPA s 2015 scholarship for honours
DOWNLOAD OR READ : PRELIMINARY AMPHIBIAN AND REPTILE SURVEY OF THE SIOUX DISTRICT OF THE CUSTER NATIONAL FOREST PDF EBOOK EPUB MOBI
 DOWNLOAD OR READ : PRELIMINARY AMPHIBIAN AND REPTILE SURVEY OF THE SIOUX DISTRICT OF THE CUSTER NATIONAL FOREST PDF EBOOK EPUB MOBI Page 1 Page 2 preliminary amphibian and reptile survey of the sioux district
DOWNLOAD OR READ : PRELIMINARY AMPHIBIAN AND REPTILE SURVEY OF THE SIOUX DISTRICT OF THE CUSTER NATIONAL FOREST PDF EBOOK EPUB MOBI Page 1 Page 2 preliminary amphibian and reptile survey of the sioux district
of Veterinary and Pharmaceutical Sciences Brno, Palackeho tr. 1/3, Brno, , Czech Republic
 Biological Journal of the Linnean Society, 2016, 117, 305 321. Comparative phylogeographies of six species of hinged terrapins (Pelusios spp.) reveal discordant patterns and unexpected differentiation
Biological Journal of the Linnean Society, 2016, 117, 305 321. Comparative phylogeographies of six species of hinged terrapins (Pelusios spp.) reveal discordant patterns and unexpected differentiation
Biology 164 Laboratory
 Biology 164 Laboratory CATLAB: Computer Model for Inheritance of Coat and Tail Characteristics in Domestic Cats (Based on simulation developed by Judith Kinnear, University of Sydney, NSW, Australia) Introduction
Biology 164 Laboratory CATLAB: Computer Model for Inheritance of Coat and Tail Characteristics in Domestic Cats (Based on simulation developed by Judith Kinnear, University of Sydney, NSW, Australia) Introduction
7.013 Spring 2005 Problem Set 2
 MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel NAME TA 7.013 Spring 2005 Problem Set 2 FRIDAY February
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel NAME TA 7.013 Spring 2005 Problem Set 2 FRIDAY February
Dynamic evolution of venom proteins in squamate reptiles. Nicholas R. Casewell, Gavin A. Huttley and Wolfgang Wüster
 Dynamic evolution of venom proteins in squamate reptiles Nicholas R. Casewell, Gavin A. Huttley and Wolfgang Wüster Supplementary Information Supplementary Figure S1. Phylogeny of the Toxicofera and evolution
Dynamic evolution of venom proteins in squamate reptiles Nicholas R. Casewell, Gavin A. Huttley and Wolfgang Wüster Supplementary Information Supplementary Figure S1. Phylogeny of the Toxicofera and evolution
What are taxonomy, classification, and systematics?
 Topic 2: Comparative Method o Taxonomy, classification, systematics o Importance of phylogenies o A closer look at systematics o Some key concepts o Parts of a cladogram o Groups and characters o Homology
Topic 2: Comparative Method o Taxonomy, classification, systematics o Importance of phylogenies o A closer look at systematics o Some key concepts o Parts of a cladogram o Groups and characters o Homology
Inheritance of Livershunt in Irish Wolfhounds By Maura Lyons PhD
 Inheritance of Livershunt in Irish Wolfhounds By Maura Lyons PhD Glossary Gene = A piece of DNA that provides the 'recipe' for an enzyme or a protein. Gene locus = The position of a gene on a chromosome.
Inheritance of Livershunt in Irish Wolfhounds By Maura Lyons PhD Glossary Gene = A piece of DNA that provides the 'recipe' for an enzyme or a protein. Gene locus = The position of a gene on a chromosome.
Geo 302D: Age of Dinosaurs LAB 4: Systematics Part 1
 Geo 302D: Age of Dinosaurs LAB 4: Systematics Part 1 Systematics is the comparative study of biological diversity with the intent of determining the relationships between organisms. Humankind has always
Geo 302D: Age of Dinosaurs LAB 4: Systematics Part 1 Systematics is the comparative study of biological diversity with the intent of determining the relationships between organisms. Humankind has always
Genetics #2. Polyallelic Traits. Genetics can be very complicated.
 Genetics #2 Genetics can be very complicated. Polyallelic Traits When a trait is caused by more than two alleles in a population. An individual still only inherits two alleles for the trait one from each
Genetics #2 Genetics can be very complicated. Polyallelic Traits When a trait is caused by more than two alleles in a population. An individual still only inherits two alleles for the trait one from each
1 EEB 2245/2245W Spring 2014: exercises working with phylogenetic trees and characters
 1 EEB 2245/2245W Spring 2014: exercises working with phylogenetic trees and characters 1. Answer questions a through i below using the tree provided below. a. The sister group of J. K b. The sister group
1 EEB 2245/2245W Spring 2014: exercises working with phylogenetic trees and characters 1. Answer questions a through i below using the tree provided below. a. The sister group of J. K b. The sister group
Bayesian Analysis of Population Mixture and Admixture
 Bayesian Analysis of Population Mixture and Admixture Eric C. Anderson Interdisciplinary Program in Quantitative Ecology and Resource Management University of Washington, Seattle, WA, USA Jonathan K. Pritchard
Bayesian Analysis of Population Mixture and Admixture Eric C. Anderson Interdisciplinary Program in Quantitative Ecology and Resource Management University of Washington, Seattle, WA, USA Jonathan K. Pritchard
Introduction Histories and Population Genetics of the Nile Monitor (Varanus niloticus) and Argentine Black-and-White Tegu (Salvator merianae) in
 Introduction Histories and Population Genetics of the Nile Monitor (Varanus niloticus) and Argentine Black-and-White Tegu (Salvator merianae) in Florida JARED WOOD, STEPHANIE DOWELL, TODD CAMPBELL, ROBERT
Introduction Histories and Population Genetics of the Nile Monitor (Varanus niloticus) and Argentine Black-and-White Tegu (Salvator merianae) in Florida JARED WOOD, STEPHANIE DOWELL, TODD CAMPBELL, ROBERT
Do the traits of organisms provide evidence for evolution?
 PhyloStrat Tutorial Do the traits of organisms provide evidence for evolution? Consider two hypotheses about where Earth s organisms came from. The first hypothesis is from John Ray, an influential British
PhyloStrat Tutorial Do the traits of organisms provide evidence for evolution? Consider two hypotheses about where Earth s organisms came from. The first hypothesis is from John Ray, an influential British
INHERITANCE OF BODY WEIGHT IN DOMESTIC FOWL. Single Comb White Leghorn breeds of fowl and in their hybrids.
 440 GENETICS: N. F. WATERS PROC. N. A. S. and genetical behavior of this form is not incompatible with the segmental interchange theory of circle formation in Oenothera. Summary.-It is impossible for the
440 GENETICS: N. F. WATERS PROC. N. A. S. and genetical behavior of this form is not incompatible with the segmental interchange theory of circle formation in Oenothera. Summary.-It is impossible for the
Selection, Recombination and History in a Parasitic Flatworm (Echinococcus) Inferred from Nucleotide Sequences
 Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 93(5): 695-702, Sep./Oct. 1998 Selection, Recombination and History in a Parasitic Flatworm (Echinococcus) Inferred from Nucleotide Sequences KL Haag, AM Araújo,
Mem Inst Oswaldo Cruz, Rio de Janeiro, Vol. 93(5): 695-702, Sep./Oct. 1998 Selection, Recombination and History in a Parasitic Flatworm (Echinococcus) Inferred from Nucleotide Sequences KL Haag, AM Araújo,
COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST
 COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST In this laboratory investigation, you will use BLAST to compare several genes, and then use the information to construct a cladogram.
COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST In this laboratory investigation, you will use BLAST to compare several genes, and then use the information to construct a cladogram.
Comparing DNA Sequence to Understand
 Comparing DNA Sequence to Understand Evolutionary Relationships with BLAST Name: Big Idea 1: Evolution Pre-Reading In order to understand the purposes and learning objectives of this investigation, you
Comparing DNA Sequence to Understand Evolutionary Relationships with BLAST Name: Big Idea 1: Evolution Pre-Reading In order to understand the purposes and learning objectives of this investigation, you
Seed color is either. that Studies Heredity. = Any Characteristic that can be passed from parents to offspring
 Class Notes Genetic Definitions Trait = Any Characteristic that can be passed from parents to offspring Heredity The passing of traits from parent to offspring - Blood Type - Color of our Hair - Round
Class Notes Genetic Definitions Trait = Any Characteristic that can be passed from parents to offspring Heredity The passing of traits from parent to offspring - Blood Type - Color of our Hair - Round
Natural history of Xenosaurus phalaroanthereon (Squamata, Xenosauridae), a Knob-scaled Lizard from Oaxaca, Mexico
 Natural history of Xenosaurus phalaroanthereon (Squamata, Xenosauridae), a Knob-scaled Lizard from Oaxaca, Mexico Julio A. Lemos-Espinal 1 and Geoffrey R. Smith Phyllomedusa 4():133-137, 005 005 Departamento
Natural history of Xenosaurus phalaroanthereon (Squamata, Xenosauridae), a Knob-scaled Lizard from Oaxaca, Mexico Julio A. Lemos-Espinal 1 and Geoffrey R. Smith Phyllomedusa 4():133-137, 005 005 Departamento
Virtual Lab: Sex-Linked Traits Worksheet. 1. Please make sure you have read through all of the information in the
 Virtual Lab: Sex-Linked Traits Worksheet 1. Please make sure you have read through all of the information in the Questions and Information areas. If you come upon terms that are unfamiliar to you, please
Virtual Lab: Sex-Linked Traits Worksheet 1. Please make sure you have read through all of the information in the Questions and Information areas. If you come upon terms that are unfamiliar to you, please
Conservation genomics of the highly endangered Red Siskin
 Conservation genomics of the highly endangered Red Siskin Haw Chuan Lim Dept of Vertebrate Zoology & Center for Conservation Genomics Smithsonian Institution Brian Coyle Project Coordinator, Red Siskin
Conservation genomics of the highly endangered Red Siskin Haw Chuan Lim Dept of Vertebrate Zoology & Center for Conservation Genomics Smithsonian Institution Brian Coyle Project Coordinator, Red Siskin
Testing Species Boundaries in an Ancient Species Complex with Deep Phylogeographic History: Genus Xantusia (Squamata: Xantusiidae)
 vol. 164, no. 3 the american naturalist september 2004 Testing Species Boundaries in an Ancient Species Complex with Deep Phylogeographic History: Genus Xantusia (Squamata: Xantusiidae) Elizabeth A. Sinclair,
vol. 164, no. 3 the american naturalist september 2004 Testing Species Boundaries in an Ancient Species Complex with Deep Phylogeographic History: Genus Xantusia (Squamata: Xantusiidae) Elizabeth A. Sinclair,
Systematics of the Lizard Family Pygopodidae with Implications for the Diversification of Australian Temperate Biotas
 Syst. Biol. 52(6):757 780, 2003 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150390250974 Systematics of the Lizard Family Pygopodidae with Implications
Syst. Biol. 52(6):757 780, 2003 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150390250974 Systematics of the Lizard Family Pygopodidae with Implications
Horned lizard (Phrynosoma) phylogeny inferred from mitochondrial genes and morphological characters: understanding conflicts using multiple approaches
 Molecular Phylogenetics and Evolution xxx (2004) xxx xxx MOLECULAR PHYLOGENETICS AND EVOLUTION www.elsevier.com/locate/ympev Horned lizard (Phrynosoma) phylogeny inferred from mitochondrial genes and morphological
Molecular Phylogenetics and Evolution xxx (2004) xxx xxx MOLECULAR PHYLOGENETICS AND EVOLUTION www.elsevier.com/locate/ympev Horned lizard (Phrynosoma) phylogeny inferred from mitochondrial genes and morphological
Evolution of Agamidae. species spanning Asia, Africa, and Australia. Archeological specimens and other data
 Evolution of Agamidae Jeff Blackburn Biology 303 Term Paper 11-14-2003 Agamidae is a family of squamates, including 53 genera and over 300 extant species spanning Asia, Africa, and Australia. Archeological
Evolution of Agamidae Jeff Blackburn Biology 303 Term Paper 11-14-2003 Agamidae is a family of squamates, including 53 genera and over 300 extant species spanning Asia, Africa, and Australia. Archeological
These small issues are easily addressed by small changes in wording, and should in no way delay publication of this first- rate paper.
 Reviewers' comments: Reviewer #1 (Remarks to the Author): This paper reports on a highly significant discovery and associated analysis that are likely to be of broad interest to the scientific community.
Reviewers' comments: Reviewer #1 (Remarks to the Author): This paper reports on a highly significant discovery and associated analysis that are likely to be of broad interest to the scientific community.
Name: Date: Hour: Fill out the following character matrix. Mark an X if an organism has the trait.
 Name: Date: Hour: CLADOGRAM ANALYSIS What is a cladogram? It is a diagram that depicts evolutionary relationships among groups. It is based on PHYLOGENY, which is the study of evolutionary relationships.
Name: Date: Hour: CLADOGRAM ANALYSIS What is a cladogram? It is a diagram that depicts evolutionary relationships among groups. It is based on PHYLOGENY, which is the study of evolutionary relationships.
EVIDENCE FOR PARALLEL ECOLOGICAL SPECIATION IN SCINCID LIZARDS OF THE EUMECES SKILTONIANUS SPECIES GROUP (SQUAMATA: SCINCIDAE)
 Evolution, 56(7), 2002, pp. 1498 1513 EVIDENCE FOR PARALLEL ECOLOGICAL SPECIATION IN SCINCID LIZARDS OF THE EUMECES SKILTONIANUS SPECIES GROUP (SQUAMATA: SCINCIDAE) JONATHAN Q. RICHMOND 1,2 AND TOD W.
Evolution, 56(7), 2002, pp. 1498 1513 EVIDENCE FOR PARALLEL ECOLOGICAL SPECIATION IN SCINCID LIZARDS OF THE EUMECES SKILTONIANUS SPECIES GROUP (SQUAMATA: SCINCIDAE) JONATHAN Q. RICHMOND 1,2 AND TOD W.
Rostral Horn Evolution Among Agamid Lizards of the Genus. Ceratophora Endemic to Sri Lanka
 Rostral Horn Evolution Among Agamid Lizards of the Genus Ceratophora Endemic to Sri Lanka James A. Schulte II 1, J. Robert Macey 2, Rohan Pethiyagoda 3, Allan Larson 1 1 Department of Biology, Box 1137,
Rostral Horn Evolution Among Agamid Lizards of the Genus Ceratophora Endemic to Sri Lanka James A. Schulte II 1, J. Robert Macey 2, Rohan Pethiyagoda 3, Allan Larson 1 1 Department of Biology, Box 1137,
Introduction to Cladistic Analysis
 3.0 Copyright 2008 by Department of Integrative Biology, University of California-Berkeley Introduction to Cladistic Analysis tunicate lamprey Cladoselache trout lungfish frog four jaws swimbladder or
3.0 Copyright 2008 by Department of Integrative Biology, University of California-Berkeley Introduction to Cladistic Analysis tunicate lamprey Cladoselache trout lungfish frog four jaws swimbladder or
Interspecific hybridization between Mauremys reevesii and Mauremys sinensis: Evidence from morphology and DNA sequence data
 African Journal of Biotechnology Vol. 10(35), pp. 6716-6724, 13 July, 2011 Available online at http://www.academicjournals.org/ajb DOI: 10.5897/AJB11.063 ISSN 1684 5315 2011 Academic Journals Full Length
African Journal of Biotechnology Vol. 10(35), pp. 6716-6724, 13 July, 2011 Available online at http://www.academicjournals.org/ajb DOI: 10.5897/AJB11.063 ISSN 1684 5315 2011 Academic Journals Full Length
Biology 2108 Laboratory Exercises: Variation in Natural Systems. LABORATORY 2 Evolution: Genetic Variation within Species
 Biology 2108 Laboratory Exercises: Variation in Natural Systems Ed Bostick Don Davis Marcus C. Davis Joe Dirnberger Bill Ensign Ben Golden Lynelle Golden Paula Jackson Ron Matson R.C. Paul Pam Rhyne Gail
Biology 2108 Laboratory Exercises: Variation in Natural Systems Ed Bostick Don Davis Marcus C. Davis Joe Dirnberger Bill Ensign Ben Golden Lynelle Golden Paula Jackson Ron Matson R.C. Paul Pam Rhyne Gail
Are Turtles Diapsid Reptiles?
 Are Turtles Diapsid Reptiles? Jack K. Horner P.O. Box 266 Los Alamos NM 87544 USA BIOCOMP 2013 Abstract It has been argued that, based on a neighbor-joining analysis of a broad set of fossil reptile morphological
Are Turtles Diapsid Reptiles? Jack K. Horner P.O. Box 266 Los Alamos NM 87544 USA BIOCOMP 2013 Abstract It has been argued that, based on a neighbor-joining analysis of a broad set of fossil reptile morphological
Bioinformatics: Investigating Molecular/Biochemical Evidence for Evolution
 Bioinformatics: Investigating Molecular/Biochemical Evidence for Evolution Background How does an evolutionary biologist decide how closely related two different species are? The simplest way is to compare
Bioinformatics: Investigating Molecular/Biochemical Evidence for Evolution Background How does an evolutionary biologist decide how closely related two different species are? The simplest way is to compare
1. Describe the series of steps that you would perform to isolate arginine-requiring mutants from a wild-type haploid yeast strain.
 1. Describe the series of steps that you would perform to isolate arginine-requiring mutants from a wild-type haploid yeast strain. i. mutagenize yeast cells. ii. plate out mutagenized yeast cells on complete
1. Describe the series of steps that you would perform to isolate arginine-requiring mutants from a wild-type haploid yeast strain. i. mutagenize yeast cells. ii. plate out mutagenized yeast cells on complete
WILDCAT HYBRID SCORING FOR CONSERVATION BREEDING UNDER THE SCOTTISH WILDCAT CONSERVATION ACTION PLAN. Dr Helen Senn, Dr Rob Ogden
 WILDCAT HYBRID SCORING FOR CONSERVATION BREEDING UNDER THE SCOTTISH WILDCAT CONSERVATION ACTION PLAN Dr Helen Senn, Dr Rob Ogden Wildcat Hybrid Scoring For Conservation Breeding under the Scottish Wildcat
WILDCAT HYBRID SCORING FOR CONSERVATION BREEDING UNDER THE SCOTTISH WILDCAT CONSERVATION ACTION PLAN Dr Helen Senn, Dr Rob Ogden Wildcat Hybrid Scoring For Conservation Breeding under the Scottish Wildcat
RICHARD D. DURTSCHE B.S. Biology, B.A. Chemistry. University of Minnesota, Duluth
 RICHARD D. DURTSCHE Department of Biological Sciences Tel: work (859) 572-6637 and Center for Natural Sciences and Mathematics home (513) 528-5290 Northern Kentucky University FAX (859) 572-5639 Highland
RICHARD D. DURTSCHE Department of Biological Sciences Tel: work (859) 572-6637 and Center for Natural Sciences and Mathematics home (513) 528-5290 Northern Kentucky University FAX (859) 572-5639 Highland
Evolution in dogs. Megan Elmore CS374 11/16/2010. (thanks to Dan Newburger for many slides' content)
 Evolution in dogs Megan Elmore CS374 11/16/2010 (thanks to Dan Newburger for many slides' content) Papers for today Vonholdt BM et al (2010). Genome-wide SNP and haplotype analyses reveal a rich history
Evolution in dogs Megan Elmore CS374 11/16/2010 (thanks to Dan Newburger for many slides' content) Papers for today Vonholdt BM et al (2010). Genome-wide SNP and haplotype analyses reveal a rich history
Molecular Phylogenetics of Squamata: The Position of Snakes, Amphisbaenians, and Dibamids, and the Root of the Squamate Tree
 Syst. Biol. 53(5):735 757, 2004 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150490522340 Molecular Phylogenetics of Squamata: The Position of
Syst. Biol. 53(5):735 757, 2004 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150490522340 Molecular Phylogenetics of Squamata: The Position of
Marsupial Mole. Notoryctes species. Amy Mutton Zoologist Species and Communities Branch Science and Conservation Division
 Marsupial Mole Notoryctes species Amy Mutton Zoologist Species and Communities Branch Science and Conservation Division Scientific classification Kingdom: Phylum: Class: Infraclass: Order: Family: Animalia
Marsupial Mole Notoryctes species Amy Mutton Zoologist Species and Communities Branch Science and Conservation Division Scientific classification Kingdom: Phylum: Class: Infraclass: Order: Family: Animalia
Mr. Bouchard Summer Assignment AP Biology. Name: Block: Score: / 20. Topic: Chemistry Review and Evolution Intro Packet Due: 9/4/18
 Name: Block: Score: / 20 Topic: Chemistry Review and Evolution Intro Packet Due: 9/4/18 Week Schedule Monday Tuesday Wednesday Thursday Friday In class discussion/activity NONE NONE NONE Syllabus and Course
Name: Block: Score: / 20 Topic: Chemistry Review and Evolution Intro Packet Due: 9/4/18 Week Schedule Monday Tuesday Wednesday Thursday Friday In class discussion/activity NONE NONE NONE Syllabus and Course
Student Exploration: Mouse Genetics (One Trait)
 Name: Date: Student Exploration: Mouse Genetics (One Trait) Vocabulary: allele, DNA, dominant allele, gene, genotype, heredity, heterozygous, homozygous, hybrid, inheritance, phenotype, Punnett square,
Name: Date: Student Exploration: Mouse Genetics (One Trait) Vocabulary: allele, DNA, dominant allele, gene, genotype, heredity, heterozygous, homozygous, hybrid, inheritance, phenotype, Punnett square,
Quiz Flip side of tree creation: EXTINCTION. Knock-on effects (Crooks & Soule, '99)
 Flip side of tree creation: EXTINCTION Quiz 2 1141 1. The Jukes-Cantor model is below. What does the term µt represent? 2. How many ways can you root an unrooted tree with 5 edges? Include a drawing. 3.
Flip side of tree creation: EXTINCTION Quiz 2 1141 1. The Jukes-Cantor model is below. What does the term µt represent? 2. How many ways can you root an unrooted tree with 5 edges? Include a drawing. 3.
Evolution of Birds. Summary:
 Oregon State Standards OR Science 7.1, 7.2, 7.3, 7.3S.1, 7.3S.2 8.1, 8.2, 8.2L.1, 8.3, 8.3S.1, 8.3S.2 H.1, H.2, H.2L.4, H.2L.5, H.3, H.3S.1, H.3S.2, H.3S.3 Summary: Students create phylogenetic trees to
Oregon State Standards OR Science 7.1, 7.2, 7.3, 7.3S.1, 7.3S.2 8.1, 8.2, 8.2L.1, 8.3, 8.3S.1, 8.3S.2 H.1, H.2, H.2L.4, H.2L.5, H.3, H.3S.1, H.3S.2, H.3S.3 Summary: Students create phylogenetic trees to
