Blood-feeding, susceptibility to infection with Schmallenberg virus and phylogenetics of Culicoides (Diptera: Ceratopogonidae) from the United Kingdom
|
|
|
- Ashlee Webb
- 5 years ago
- Views:
Transcription
1 Barber et al. Parasites & Vectors (2018) 11:116 DOI /s x RESEARCH Blood-feeding, susceptibility to infection with Schmallenberg virus and phylogenetics of Culicoides (Diptera: Ceratopogonidae) from the United Kingdom James Barber 1, Lara E. Harrup 1, Rhiannon Silk 1, Eva Veronesi 1,2, Simon Gubbins 1, Katarzyna Bachanek-Bankowska 1 and Simon Carpenter 1* Open Access Abstract Background: Culicoides biting midges (Diptera: Ceratopogonidae) are responsible for the biological transmission of internationally important arboviruses of livestock. In 2011, a novel Orthobunyavirus was discovered in northern Europe causing congenital malformations and abortions in ruminants. From field studies, Culicoides were implicated in the transmission of this virus which was subsequently named Schmallenberg virus (SBV), but to date no assessment of susceptibility to infection of field populations under standardised laboratory conditions has been carried out. We assessed the influence of membrane type (chick skin, collagen, Parafilm M ) when offered in conjunction with an artificial blood-feeding system (Hemotek, UK) on field-collected Culicoides blood-feeding rates. Susceptibility to infection with SBV following blood-feeding on an SBV-blood suspension provided via either (i) the Hemotek system or via (ii) a saturated cotton wool pledglet was then compared. Schmallenberg virus susceptibility was defined by RT-qPCR of RNA extractions of head homogenates and related to Culicoides species and haplotype identifications based on the DNA barcode region of the mitochondrial cytochrome c oxidase1(cox1) gene. Results: Culicoides blood-feeding rates were low across all membrane types tested (7.5% chick skin, 0.0% for collagen, 4.4% Parafilm M, with 6029 female Culicoides being offered a blood meal in total). Susceptibility to infection with SBV through membrane blood-feeding (8 of 109 individuals tested) and pledglet blood-feeding (1 of 94 individuals tested) was demonstrated for the Obsoletus complex, with both C. obsoletus (Meigen) and C. scoticus Downes & Kettle susceptible to infection with SBV through oral feeding. Potential evidence of cryptic species within UK populations was found for the Obsoletus complex in phylogenetic analyses of cox1 DNA barcodes of 74 individuals assessed from a single field-site. Conclusions: Methods described in this study provide the means to blood-feed Palaearctic Culicoides for vector competence studies and colonisation attempts. Susceptibility to SBV infection was 7.3% for membrane-fed members of the subgenus Avaritia and 1.1% for pledglet-fed. Both C. obsoletus and C. scoticus were confirmed as being susceptible to infection with SBV, with potential evidence of cryptic species within UK Obsoletus complex specimens, however the implications of cryptic diversity in the Obsoletus complex on arbovirus transmission remains unknown. Keywords: Vector competence, Biting midges, Arbovirus, SBV, Orthobunyavirus, DNA barcode, cox1 * Correspondence: simon.carpenter@pirbright.ac.uk; simon.carpenter@pirbright.ac.uk Equal contributors 1 Vector-borne Viral Disease Programme, The Pirbright Institute, Pirbright, Surrey, UK Full list of author information is available at the end of the article The Author(s) Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License ( which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver ( applies to the data made available in this article, unless otherwise stated.
2 Barber et al. Parasites & Vectors (2018) 11:116 Page 2 of 13 Background In 2011, a novel Orthobunyavirus, provisionally named Schmallenberg virus (SBV), was discovered in Germany and was subsequently found to cause foetal abnormalities and abortions in ruminants [1 3]. To date, no convincing route for the incursion of SBV into Europe has been provided, echoing the outbreak of bluetongue virus (BTV) serotype 8 that occurred in the same region 5 years prior to this event [4]. Subsequently, surveys based upon detection of SBV antibodies, viral RNA and clinical disease in livestock demonstrated that the virus spread rapidly across the northwestern region of Europe during [5 8]. Countries in the region then reported significant reductions in additional clinical cases and seroconversion to SBV infection during [5, 9], driven by a lack of naïve hosts. Since these studies, recirculation of the virus in northwestern Europe has occurred [10], but the majority of monitoring surveys have been scaled back following reductions in clinical cases of disease and the deployment of an effective vaccine [11]. Culicoides biting midges (Diptera: Ceratopogonidae) were suspected as the primary biological vectors of SBV prior to direct study of transmission in the field, due to their involvement in the BTV epidemic in northwestern Europe 5 years earlier [12], and the close phylogenetic relationship of SBV with other Culicoides-borne orthobunyaviruses [1, 13]. This hypothesis was confirmed by a series of field-based trials in Belgium, the Netherlands and France that detected significant quantities of SBV RNA in Culicoides collected in close proximity to livestock [14 18], while failing to detect virus in mosquitoes [19]. Techniques to assess the probability of field transmission of SBV by Culicoides and mosquitoes were standardised using colony lines of Culicoides sonorensis Wirth & Jones and Culicoides nubeculosus (Meigen) infected using artificial membrane-based techniques [20, 21]. Within northwestern Europe, the Culicoides fauna on farms and stables are dominated by species classified within the subgenus Avaritia [18, 22, 23]. Until recently four species within this subgenus have been identified within this region and are commonly referred to as the Obsoletus group, despite a lack of monophyly: Culicoides obsoletus (Meigen) and Culicoides scoticus Downes & Kettle; Culicoides dewulfi Goetghebuer and Culicoides chiopterus (Meigen) [24 26]. Several additional cryptic species have also recently been proposed whose taxonomic status, prevalence, abundance and involvement in transmission of arboviruses remains uncertain [27 29]. To date, these cryptic species have not been identified in the UK, despite a study that examined 79 individuals of the subgenus Avaritia using a 472 bp region of the mitochondrial cytochrome c oxidase subunit 1 gene (cox1) across ten geographically disparate sampling points [25]. All four established members of Culicoides (Avaritia)in northwestern Europe have been implicated in transmission of SBV. Schmallenberg virus RNA has been detected in homogenates of heads removed from field-collected C. obsoletus, C. scoticus and C. chiopterus individuals in the Netherlands [15], and additionally within heads of C. dewulfi collected in Belgium [17]. A key research limitation, however, is that techniques have not been developed to artificially blood-feed field-collected species of Palaearctic Culicoides through membranes under conditions of biological containment [30, 31]. This not only limits studies attempting to standardise population susceptibility to infection across the region, but also prevents the establishment of colony and cell lines of these species as a resource [32]. In this study we examined methods of blood-feeding species of field-collected Culicoides in the UK to provide a standard for comparative studies of vector competence across Europe. We then examined vector competence in a single population of Culicoides collected at a stable in Surrey, UK. To our knowledge, for the first time, we link standard detection of disseminated infections with haplotype-based identification based on Sanger-sequencing of the DNA barcode region of the mitochondrial cytochrome c oxidase subunit 1 (cox1) gene. This not only provides preliminary data concerning susceptibility rates to SBV infection in known vector species of Culicoides in northwestern Europe, but also provides additional data concerning the complement of species and cox1 haplotypes that are present in the UK, with consequences for epidemiological studies of arbovirus transmission and epidemiology. Methods Study site and collections Specimens of Culicoides were collected at a site in close proximity to horses at a stable in Surrey ( N; W) during 2012 and 2013 using two 8 W ultraviolet Onderstepoort Veterinary Institute (OVI) light-suction traps (Agricultural Research Council - Onderstepoort Veterinary Institute, Pretoria, South Africa) [33]. Collections were made overnight in a 500 ml plastic beaker partially filled with damp paper hand-towel in an attempt to minimise the effects of desiccation on collected Culicoides. Following collection, collected insects were allowed to emerge from the beaker against a glass plane in front of natural daylight allowing selected aspiration of free-flying Culicoides for use during later experiments. These were transferred to netted, cylindrical, card 64 mm diameter pillboxes (Watkins and Doncaster, Stainton, UK) for use in later experiments. Effect of membrane type on blood-feeding rates All blood-feeding studies were conducted in 2012 using an artificial feeding system (Hemotek), with each feeder
3 Barber et al. Parasites & Vectors (2018) 11:116 Page 3 of 13 unit calibrated to warm the blood-meal to 37 C. Prior to being offered a blood-meal, all Culicoides were incubated for 96 h following collection, at 25 ± 1 C and 50% relative humidity (RH) with ad libitum access to 10% (w/v) sucrose solution (Sigma-Aldrich, Gillingham, UK) provided via a saturated cotton wool pledglet placed on the netted tops of pillboxes. Pledglets were removed 24 h prior to offering Culicoides a blood meal. Three membrane types were compared, used in conjunction with an artificial feeding system (Hemotek). Blood-meal reservoirs containing approximately 3 ml of defibrinated equine blood (TCS Biosciences, Botolph Claydon, UK) were covered with either a (i) stretched skins taken from one-day old-chicks; (ii) collagen membrane (Hemotek); or (iii) stretched Parafilm M membrane (Bemis Company Inc., Neenah, WI, USA). Each membrane type was assessed in duplicate for each round of blood-feeding with membrane types randomly allocated two of the six feeders units per blood-feeding round. A total of nine days of duplicate replicates were completed for each membrane type. Blood meals were offered for a total of 45 min, after which the contents of the pillboxes were killed via prolonged exposure to cold and Culicoides were identified to species or subgenus level according to their morphology with reference of appropriate keys [24]. In addition specimens were classified according to abdominal pigmentation (unpigmented or pigmented); blood-fed or gravid in the case of females [34, 35], or as males. Blood-feeding success rates were calculated using only unpigmented, pigmented and blood-fed females. The effect of membrane type and date of blood-feeding on the number of blood-fed Culicoides was assessed using a Kruskal- Wallis test. Where the Kruskal-Wallis test was significant (P < 0.05), differences between factor levels were explored using pairwise Wilcoxon rank-sum tests. Vector competence All vector competence studies were conducted in August-October The SBV strain used for studies of infection was provided by IZS Teramo from an isolation originally made by the Friedrich-Loeffler-Institute, Isle of Riems, Germany [1]. This SBV strain had been passaged once through a C. sonorensis derived cell line and four times through a baby hamster kidney (BHK-21) cell line and then adjusted to a titre of 10 6 tissue culture infectious dose 50 (TCID 50 ) using defibrinated equine blood (TCS Bioscience). All blood-feeding and sorting of Culicoides were carried out in a biosecure glove-box. Blood-virus mixes were offered using either (i) an artificial feeding system (Hemotek) as described above with a stretched Parafilm M membrane (Bemis Company Inc.), or via (ii) a saturated cotton wool pledglet placed directly on top of the net of the pillboxes [31]. Blood meals were offered for a total of 45 min, after which blood-fed Culicoides were selected under light CO 2 anaesthesia and then incubated in netted, cylindrical, card 64 mm diameter pillboxes (Watkins and Doncaster) for eight days at 25 C with ad libitum access to 10% (w/v) sucrose solution (Sigma-Aldrich) provided via a cotton wool pledglet placed on top of the netted pillboxes. Culicoides surviving the incubation period were then selected under CO 2 anaesthesia and transferred to 1.5 ml Eppendorf tubes containing 70% ethanol and stored at 4 C prior to further analysis. Specimens were individually removed from storage in 70% ethanol and decapitated using a sterile needle (Monoject hypodermic needle, 18g 1.5; Covidien, Minneapolis, MN, USA). Heads were then transferred individually to 1.5 ml microcentrifuge tubes containing 100 μl Schneider s Drosophila Media (Gibco, Paisley, UK) and homogenised using disposable polypropylene pestles (Sigma-Aldrich). The remaining abdomen and thorax of Table 1 Comparison of membrane type with an artificial feeding system (Hemotek, UK) on Culicoides blood-feeding rate (blood-feeding rate with range and total number offered a blood meal shown in parentheses). Data exclude gravid females and males exposed to blood-feeding system Culicoides spp./ Species complex Blood-feeding rate (%) Chick skin Collagen Parafilm M Obsoletus complex (n = 4270) 8.0 (0 16.0; n = 1384) 0 (n = 1331) 5.0 (0 32.0; n = 1555) C. dewulfi Goetghebuer (n = 121) 7.0 (0 33.0, n = 27) 0 (n = 50) 0 (n = 44) C. chiopterus (Meigen) (n =2) 0(n =1) C. pulicaris (L.) (n = 34) 0.03 (0 33.0; n = 34) C. punctatus (Meigen) (n = 85) 0 (n = 37) 0 (n = 19) 0.10 ( ; n = 29) C. impunctatus Goetghebuer (n =4) 0(n =1) 0(n =2) 0(n =1) C. achrayi (Kettle & Lawson) (n = 255) 8.0 (0 33.0; n = 85) 0 (n = 87) 2.0 (0 25.0; n = 83) C. festivipennis Kieffer (n = 43) 30.8 ( ; n = 13) 0 (n =6) 0(n = 24) Other Culicoides spp. (n =4) 0(n =4) Total blood-fed (%) 119 (7.9) 0 (0) 76 (4.3)
4 Barber et al. Parasites & Vectors (2018) 11:116 Page 4 of 13 Table 2 Susceptibility to infection with Schmallenberg virus for Culicoides (Avaritia) collected at a single site in the UK Subgenus Avaritia Feeding method Membrane a Pledglet a 109 (13) 94 (2) C. pulicaris (L.) 2 (2) 4 (0) C. punctatus (Meigen) 4 (0) 6 (0) C. achrayi (Kettle & Lawson) 23 (0) C. festivipennis Kieffer 9 (0) Total 147 (15) 104 (2) a n with total with a quantification cycle (C q ) 40 in parenthesis each individual was transferred to a collection microtube (Qiagen, Manchester, UK) containing 100 μl of Schneider s Drosophila Media and a 3 mm stainless steel bead (Dejay Distribution Ltd., Launceston, UK) and homogenised for 1 min at 25 hz using a TissueLyser (Qiagen). Schmallenberg virus detection Total nucleic acid was extracted from the homogenates of Culicoides heads using a Universal Biorobot (Qiagen) with a QIAamp All Nucleic Acid MDx Kit (Qiagen) following the manufacturer s recommended instructions. Schmallenberg virus RNA in the resultant extractions was the assessed using a semi-quantitative RT-qPCR targeting the S segment of the genome [1, 20]. A C q cut-off value of < 35 was used to define SBV infection [15]. cox1 DNA barcode assay Amplification of a 658 bp fragment of the DNA barcoding region of the mitochondrial cox1 gene[36] was achieved by polymerase chain reaction (PCR). Reactions were performed in a total volume of 25 μl consistingof13.65μl nuclease-free water (Qiagen), 2.5 μl 10 PCR Buffer (Life Technologies, Paisley, UK), 0.75 μl 50 mm MgCl 2 (Life Technologies), 0.5 μl 10 mm dntp mix (Life Technologies); 0.1 μl Platinum Taq DNA polymerase (Life Technologies), 1.25 μl of the 20 μm forward primer, 1.25 μl of the 20 μm reverse primer and 5.0 μl oftemplatedna (approximately 5 25 ng DNA) per reaction. Amplification of the DNA barcode region was initially attempted using the following primer pair: LCO1490 (5 -GTC AAC AAA TCA TAA AGA TAT TGG-3 [37]) and HCO2198 (5 - TAA ACT TCA GGG TGA CCA AAA AAT CA-3 [37]). If the DNA barcode region could not be sucessfully amplified using primers LCO1490 and HCO2198, the above assay was repeated using the following alternative primer pair: LepF1 (5 -ATT CAA CCA ATC ATA AAG ATA TTG G-3 [38]) and LepR1 (5 -TAA ACT TCT GGA Table 3 Observed quantification cycle (C q ) values for Schmallenberg virus (SBV) in field-collected Culicoides. Includes associated Culicoides species characterisation and blood-feeding method ( indicates no sequence data available; Obsoletus complex includes C. obsoletus (Meigen) and C. scoticus Downes & Kettle) Species BIN a Blood-feeding method SBV C q value Culicoides DNA barcode Sample ID GenBank ID BOLD process ID C. obsoletus (Meigen) BOLD:AAO7718 Membrane TPI:ENT:# KT CUSBV Membrane TPI:ENT:# KT CUSBV Membrane TPI:ENT:# KT CUSBV Membrane TPI:ENT:# KT CUSBV Membrane TPI:ENT:# KT CUSBV Membrane TPI:ENT:# KT CUSBV Membrane TPI:ENT:# KT CUSBV Membrane TPI:ENT:# KT CUSBV Membrane TPI:ENT:# KT CUSBV BOLD:AAM6198 Membrane TPI:ENT:# KT CUSBV C. scoticus Downes & Kettle BOLD:AAZ3985 Pledglet TPI:ENT:# KT CUSBV Pledglet TPI:ENT:# KT CUSBV Obsoletus complex b Membrane Membrane Membrane C. pulicaris (L.) b Membrane Membrane a Barcode Index Numbers (BINs) [49] assigned within the Barcode of Life Database (BOLD) [39] for specimens collected within this study: specimens TPI:ENT:# TPI:ENT:# ; TPI:ENT:# TPI:ENT:# ; TPI:ENT:# TPI:ENT:# ; TPI:ENT:# TPI:ENT:# [GenBank: KT KT186811; KT KT186857; KT KT186862; KT KT186879] b Species identification based on morphology only
5 Barber et al. Parasites & Vectors (2018) 11:116 Page 5 of 13 TGT CCA AAA AAT CA-3 [38]). Positive and negative controls for the amplification reactions were carried out at every PCR round. The PCR cycling conditions for all DNA Barcode assays were as follows: an initial denaturation step at94 Cfor2minfollowedby35cyclesof94 Cfor30s, 46 Cfor30s,72 Cfor1min,andafinalextensionstepat 72 C for 10 min. PCR products were visualised using ultraviolet light and 2% agarose gels containing SYBR Safe DNA Gel Stain (Invitrogen, Paisley, UK). Successful amplification of the cox1 DNA barcode region was indicated by the presence of a band at approximately 720 bp for both primer pair LCO1490 and HCO2198 and primer pair LepF1 and LepR1, identified by comparison with E-Gel Low Range Quantitative DNA Ladder ( bp; Life Technologies). PCR purification and cox1 sequencing Dimer formation from the primers was not observed and purification of the remaining PCR product was performed using a GFX TM PCR DNA and Gel Band Purification Kit (GE Healthcare Life Sciences, Little Chalfont, UK), following the manufacturer s recommended guidelines. The amplicons were sequenced bi-directionally using BigDye Terminator v3.1 Cycle Sequencing kit (Applied Biosystems, Paisley, UK) in a 3730 DNA Analyzer (Applied Biosystems) according to the manufacturer s instructions using the primer pairs as per the PCR amplification either HCO2198 and LCO1490, or LepF1 and LepR1. All sequences were aligned using MUSCLE [41] and the alignment quality checked using GUIDANCE [42] (100 bootstraps). All included sequences were aligned with a high degree of confidence (GUIDANCE alignment score > 0.999). jmodeltest [43, 44] v was then used to determine the most suitable DNA substitution model among the 24 models that can be implemented in MrBayes [45, 46]. The model with the lowest Bayesian information criterion (BIC) and Akaike information criterion (AIC), the Hasegawa-Kishino-Yano with gamma-distribution rates (HKY+G) [47], was considered to best describe the nucleotide substitution pattern. The phylogenetic relationships among taxa were resolved using a Bayesian inference (BI) approach using the HKY+G nucleotide substitution model rooted on the partial cox1 sequence of C. imicola Kieffer (Gen- Bank: KT307824) [48]. The BI tree was constructed using MrBayes v [46, 49] and twenty million tree generations in four chains were run, sampling every 1000th and discarding the first 25%, before constructing a 50% majority rule consensus tree reporting Bayesian posterior probabilities. Convergence was assessed using Phylogenetic analysis Electropherograms were edited and forward and reverse sequences assembled and trimmed to remove primer sequence using CodonCode Aligner v (CodonCode Aligner, Centerville, MA, USA). Corresponding specimen collection data and DNA sequences including electropherograms have been made publically available via the Barcode of Life Data System (BOLD) [39] as dataset DS-CUSBV (dx.doi.org/ /ds-cusbv), DNA sequences were also submitted to the GenBank database (accession numbers KT KT186881). Consensus sequences were compared to previously published sequences in GenBank using the standard nucleotide BLAST tool [40], in addition to comparison to as yet unreleased sequence data in the BOLD database [39] using the Barcode Identification Engine in BOLD v3. Obsoletus group cox1 sequences which overlapped the DNA barcode region [36] by at least 390 bp were obtained from both BOLD (n = 25) and Gen- Bank (n = 393) and included in the phylogenetic analysis (Additional file 1: Table S1), these sequences were selected in order to assess if the haplotypes identified within this study were concordant with those identified from other geographical regions. Fig. 1 Most parsimonious median-joining network (ε = 0) depicting phylogenetic relationships among C. obsoletus cox1 haplotypes. The size of each circle is proportional to the corresponding haplotype frequency. Circles are coloured to represent the proportion of specimens which showed a positive (red) and negative (blue) SBV response as determined by qpcr. Branch lengths are proportional to the number of nucleotide changes between haplotypes. Black circles indicate median vectors (mv) that represent hypothetical missing or unsampled ancestral haplotypes
6 Barber et al. Parasites & Vectors (2018) 11:116 Page 6 of 13 AWTY [50]. The BI trees were visualised using the packages Ape v. 3.2 [51] and Phytools v [52] implemented in R v [53]. Relationships between the observed haplotypes within the C. obsoletus specimens for which SBV vector competence data were available (n = 67) (GenBank: KT KT186878), were assessed by constructing Median-Joining networks. Roehl haplotype data files (RDF) were created with DnaSP v5.10 [54] and imported into Network v [55]. Networks were calculated with the Median-Joining algorithm [56] with equal weights for all characters, using maximum parsimony [57] post-processing. Intra- and inter-specific uncorrected percent nucleotide sequence distances, were generated using the packages Spider v [58] and Ape v. 3.2[51] implemented in R v [53]. Missing nucleotides were treated in all sequence comparisons using a pairwise deletion option. Results Effect of membrane type on blood-feeding A total of 6029 Culicoides were used in the experiments to investigate the effect of membrane type on Culicoides blood-feeding rates: 4681 specimens of the Obsoletus complex including the morphologically cryptic C. obsoletus (Meigen) and C. scoticus Downes & Kettle; 157 C. dewulfi; 4 C. chiopterus; 127 C. pulicaris (L.); 138 C. punctatus (Meigen), 4 C. impunctatus Goetghebuer; 321 C. achrayi Kettle & Lawson; 537 C. festivipennis Kieffer; 60 individuals from other Culicoides species (Additional file 2: Table S2). Excluding gravid females and males, a total of 4270 specimens of the Obsoletus complex; 121 C. dewulfi; 2 C. chiopterus; 96C. pulicaris; 85C. punctatus; 4C. impunctatus; 255 C. achrayi; 43 C. festivipennis were exposed to the membrane systems during trials (Table 1). Only bloodfeeding rates of the species of the Obsoletus complex provided sufficient sample sizes for further statistical analysis. Membrane type had a significant (Kruskal-Wallis test: χ 2 =27.1,df =2,P < ) effect on Obsoletus complex Culicoides blood-feeding rate, with both chick skin (Wilcoxon rank-sum test: W = 27, P < ) and Parafilm M (Wilcoxon rank-sum test: W =297,P < ) membrane resulting in a significantly higher bloodfeeding rate that the collagen membrane. There was no significant (Wilcoxon rank-sum test: W = 180, P = 0.57) difference in blood-feeding rate between Obsoletus complex Culicoides fed via chick skin and Parafilm M membrane. The day of blood-feeding also had no significant effect (Kruskal-Wallis test: χ 2 =2.7,df =7,P =0.91)on blood-feeding rates. Vector competence In total, 147 Culicoides survived the eight day incubation period post-feeding on the blood-virus mix through an artificial membrane; of these 13 (11.9%) of Culicoides (Avaritia) Fig. 2 Simplified Bayesian inference phylogenetic tree inferred from cox1 DNA barcode sequences with species assignments indicated. Bayesian posterior probability node support values 0.95 are shown as solid black circles at nodes, shown as black outline circles with grey fill at nodes and < 0.75 shown as black outline circles with white fill at nodes
7 Barber et al. Parasites & Vectors (2018) 11:116 Page 7 of 13 contained significant quantities of SBV RNA within their heads (C q 40) (Table 2). Using a more conservative cutoff level for detection of SBV RNA (C q 35), this number fell to 8 (7.3%). In total, 104 Culicoides were successfully fed using pledglet blood-feeding, of which 94 were from the subgenus Avaritia (Table 2). Of these, 2 (2.1%) contained significant quantities of SBV RNA in their heads (C q 40) (Tables 2, 3). Using a more conservative cut-off level for detection of SBV RNA (C q 35), this number fell to 1 (1.1%). In C. pulicaris fed using a membrane method both processed individuals tested positive for disseminated SBV RNA of which one had a C q value lower than the conservative cut-off (Table 3). Heads from no other species examined contained detectable levels of SBV RNA (Table 2). Phylogenetic analysis Full length primer truncated (658 bp) DNA barcode sequences (n = 74) were obtained from three species of the subgenus Avaritia: C. obsoletus (n = 67), C. scoticus (n =3)andC. dewulfi (n = 4). No insertions, deletions, amino acid frame shifts or stop codons were observed among these sequences, and their translations, indicating that pseudogenes were not present within the alignments. DNA barcodes were obtained from 12 of a total of 15 specimens containing SBV RNA in their heads (C q 40): ten from membrane-fed individuals and two from pledglet-fed individuals (Table 3). Ten were identified as C. obsoletus (Table 3), there was, however, no apparent association between haplotype and SBV vector competence within this species (Fig. 1). DNA from a further three specimens could not be amplified successfully and these were identified on the basis of morphology only as belonging to the Obsoletus complex (Table 3). DNA barcodes for an additional 46 membrane-fed Culicoides and 17 pledglet-fed Culicoides lacking detectable SBV RNA (C q 40) were also generated and analysed (Table 3). Phylogenetic analysis of the DNA barcodes from specimens generated within this study together with cox1 sequences obtained from GenBank and Fig. 3 Bayesian inference phylogenetic tree inferred from C. obsoletus cox1 DNA barcode sequences. Bayesian posterior probability node support values 0.95 are shown as solid black circles at nodes, shown as black outline circles with grey fill at nodes and < 0.75 shown as black outline circles with white fill at nodes. Geographical origin of specimens indicated by a coloured circle preceding the GenBank accession number for the sequence. Schmallenberg virus (SBV) vector competence indicates by colour of the GenBank accession number tip labels (red, positive; blue, negative; black, unknown). Clades shown represented in purple in Fig. 2
8 Barber et al. Parasites & Vectors (2018) 11:116 Page 8 of 13 Fig. 4 Bayesian inference phylogenetic tree inferred from C. scoticus cox1 DNA barcode sequences. Bayesian posterior probability node support values 0.95 are shown as solid black circles at nodes, shown as black outline circles with grey fill at nodes and < 0.75 shown as black outline circles with white fill at nodes. Geographical origin of specimens indicated by a coloured circle preceding the GenBank accession number for the sequence. Schmallenberg virus (SBV) vector competence indicates by colour of the GenBank accession number tip labels (red, positive; blue, negative; black, unknown). Clades shown represented in orange in Fig. 2 BOLD (Additional file 1: Table S1) indicated that the species clades represented in the BI phylogeny were concordant with morphological identifications generated using the key of Campbell & Pelham-Clinton [24] (Figs. 2, 3, 4, 5, 6). The cox1 sequences obtained from GenBank and BOLD overlapped the 658 bp DNA barcodes generated in this study by between 391 bp and 658 bp. Deep interspecific differences within the cox1 DNA barcode region were present between the majority of Culicoides species present within the phylogenetic analysis (Table 4, Figs. 2, 7; Additional file 1: Figures S1, S2). The greatest intraspecific nucleotide sequence difference was between C. dewulfi and C. chiopterus [mean 22.5% (range: %)], while the least was between C. obsoletus and C. scoticus [mean 12.2% (range: %)] (Table 4; Additional file 1: Figure S2). Sequence differences symptomatic of cryptic species diversity were, however, present within C. obsoletus (Table 4; Additional file 1: Figure S2). Intraspecific nucleotide sequence differences were observed between specimens nominally identified as C. obsoletus ranging from 0 to 12.3%. Barcode Index Number (BINs) [49] assigned within BOLD [39] for C. scoticus (BOLD:AAZ3985, n = 3) and C. dewulfi (BOLD:AAZ3991, n = 4) were concordant across all specimens sequenced within this study, however discordant BINs were observed for the C. obsoletus specimens sequenced within this study (Concordant BINs exhibit > 2% nucleotide sequence divergence [49]). This resulted in two BINs (BOLD:AAM6198, n = 5; BOLD:AA07718, n = 62) being assigned for specimens putatively identified as C. obsoletus thereby indicating the potential presence of two distinct taxa (see Table 3, dataset DS-CUSBV: dx.doi.org/ /ds-cusbv). Discussion This study has developed artificial blood-feeding techniques for Culicoides in the UK and then demonstrated susceptibility to infection with SBV in multiple haplotypes of C. obsoletus and C. scoticus under laboratory conditions. The results confirm those that identified SBV RNA in the heads of these species in close proximity to ruminants in the Netherlands [15], Belgium [17]
9 Barber et al. Parasites & Vectors (2018) 11:116 Page 9 of 13 Fig. 5 Bayesian inference phylogenetic tree inferred from C. dewulfi cox1 DNA barcode sequences. Bayesian posterior probability node support values 0.95 are shown as solid black circles at nodes, shown as black outline circles with grey fill at nodes and < 0.75 shown as black outline circles with white fill at nodes. Geographical origin of specimens indicated by a coloured circle preceding the GenBank accession number for the sequence. Schmallenberg virus (SBV) vector competence indicates by colour of the GenBank accession number tip labels (red, positive; blue, negative; black, unknown). Clade shown represented in green in Fig. 2 and France [14]. Blood-feeding of Culicoides on pledglets was also demonstrated to underestimate rates of susceptibility to infection, as found previously with BTV [30, 31]. The sequencing of the DNA barcode region (658 bp) of the cox1 gene for 67 individuals putatively identified as C. obsoletus from a single site in the UK provides potential evidence of cryptic diversity, however, how this relates to the morphologically cryptic species previously described in the Netherlands, Sweden and Switzerland [27 29, 59] remains unclear. Blood-feeding techniques developed within this study enabled the first membrane-based artificial feeding experiments using northwestern European Culicoides since the 1980s [60]. Unlike the original method, which relied upon the use of populations in close proximity to contained facilities, the technique had the advantage of including an incubation step that precluded the inclusion of naturally blood-fed individuals from traps and can be used with light-suction trap collections from a wide geographical area. While blood-feeding rates remained poor, this development is useful in creating a platform for both vector competence experimentation and wider vectorvirus interaction studies in addition to laboratory-based studies of bionomics for which little data is available for the subgenus Avaritia in northwestern Europe [32]. Further investigation of techniques to improve feeding rates is now required as provided for C. impunctatus in northwestern Europe [61, 62]. A high rate of vector competence for SBV in Culicoides is hypothesised to be at least partly responsible for driving the rapid geographic spread of the virus in Europe [14, 15, 63]. While the preliminary data provided in the current study shows that susceptibility to infection with SBV is low, significant variation in vector susceptibility to infection according to population tested has been documented for Culicoides of the subgenus Avaritia in the UK for BTV [30]. Hence, further screening of Culicoides populations utilising multiple populations across Europe would be beneficial in improving our understanding of transmission and in informing mathematical modelling studies [63, 64]. Based on analysis of the cox1 DNA barcode region, sequences identified as C. obsoletus group into two major phylogenetic clades (Figs. 2, 3). All 67 sequences of C. obsoletus specimens obtained in the study conformed to one of these two clade (Clade 1) alongside sequences obtained during previous UK-based studies [25]. Clade 1 is thought to represent the classical morphological description of C. obsoletus [24] while the second clade includes specimens which have previously been associated
10 Barber et al. Parasites & Vectors (2018) 11:116 Page 10 of 13 Fig. 6 Bayesian inference phylogenetic tree inferred from C. chiopterus cox1 DNA barcode sequences. Bayesian posterior probability node support values 0.95 are shown as solid black circles at nodes, shown as black outline circles with grey fill at nodes and < 0.75 shown as black outline circles with white fill at nodes. Geographical origin of specimens indicated by a coloured circle preceding the GenBank accession number for the sequence. Schmallenberg virus (SBV) vector competence indicates by colour of the GenBank accession number tip labels (red, positive; blue, negative; black, unknown). Clades shown represented in red in Fig. 2 with the dark form of C. obsoletus in continental Europe [28]. Clade 2a (Figs. 2, 3) groups sequences of specimens which have been previously referenced as C. obsoletus O2 [59], while Clade 2b (Figs. 2, 3) containsamixture of specimens which have been previously referenced as C. obsoletus O1, O2 and O3 [65]. In contrast, C. scoticus, C. chiopterus and C. dewulfi were all strongly supported as monophyletic clades with relatively low intraspecific sequence differences (Figs. 2, 4, 5, 6, Table 4). Table 4 Nucleotide sequence distances between UK Culicoides cox1 DNA barcodes. Uncorrected percent nucleotide sequence distances, mean with range shown in parentheses split by species including the proposed clades of C. obsoletus (see Fig. 2). Intraspecific distances are shown in bold along the diagonal, interspecific distances are shown below the diagonal. The number of specimens per species (n) is shown in brackets followed by the number of specimens originating from this study; and the number originating from GenBank in parentheses C. chiopterus C. dewulfi C. obsoletus C. obsoletus Clade 1 C. chiopterus [56 (0; 56)] 0.8 ( ) C. dewulfi [75 (4; 71)] ( ) ( ) C. obsoletus [169 (124; 145)] 14.9 ( ) C. obsoletus Clade 1 [206 (67; 139)] 14.6 ( ) C. obsoletus Clade 2a [6 (0; 6)] 16.2 ( ) C. obsoletus Clade 2b [57 (57; 0)] 14.7 ( ) C. scoticus [92 (3; 89)] 14.0 ( ) 19.3 ( ) 18.8 ( ) 21.0 ( ) 20.2 ( ) 4.1 ( ) 0.9 ( ) C. obsoletus Clade 2a 9.4 ( ) 1.7 ( ) C. obsoletus Clade 2b 11.1 ( ) 8.6 ( ) 0.6 ( ) C. scoticus 18.2 ( ) 12.2 ( ) 12.1 ( ) 11.6 (14.3) 12.1 ( ) 0.4 ( )
11 Barber et al. Parasites & Vectors (2018) 11:116 Page 11 of 13 Fig. 7 Most parsimonious median-joining network (ε = 0) depicting phylogenetic relationships cox1 haplotypes. The size of each circle is proportional to the corresponding haplotype frequency. Branch lengths are proportional to the number of nucleotide changes between haplotypes Within the UK, further assessment of cryptic species prevalence and additional multi-locus studies are required across a wide-geographic area, both to assess the robustness of current PCR multiplex assays [25, 66] used to determine species in studies and to contribute to understanding relationships between phylogenetics and vector competence for arboviruses within the subgenus Avaritia. At a regional scale, studies are required that link both northwestern and northeastern European fauna and Nearctic species of the subgenus Avaritia. Several species in this group remain entirely uncharacterised using molecular markers, e.g. C. sanguisuga (Coquillett). Such data will then make decisions whether the observed sequence diversity represents novel species or potentially species that require resurrection from synonymy permissible, e.g. C. dobyi Callot & Kremer [67]. To resolve these issues it is particularly important that specimens are collected from the species type-locality in order to confirm that subsequent specimens are in fact conspecific (see Harrup et al. [68] for review). Inaccurate species identification can have significant impacts on epidemiological investigations and control attempts when species have divergent vector competences and/or habitat and host preferences. Examination of the relative relationship between species within the subgenus Avaritia and confirmation of the monophyletic status of the subgenus, coupled with further arbovirus vector competence experiments will allow investigation of whether arbovirus vector competence is an ancestral character in the Obsoletus complex as previously proposed for the Imicola complex [69] or whether vector competence has evolved multiple times within this group. Conclusions Methods described in this study provide the means to blood-feed Palaearctic Culicoides for vector competence studies and colonisation attempts. Susceptibility to SBV infection was 7.3% for membrane-fed members of the subgenus Avaritia and 1.1% for pledglet-fed. Both C. obsoletus and C. scoticus were confirmed as being susceptible to infection with SBV, with potential evidence of cryptic species within UK Obsoletus complex specimens; however the implications of cryptic diversity in the Obsoletus complex on arbovirus transmission remains unknown. This is important as these species are the most abundant Culicoides biting midges on farms across northwestern Europe and arboviruses transmitted by them continue to emerge in this region. Additional files Additional file 1: Table S1. GenBank sequences used in the genetic analyses of Obsoletus complex of Culicoides. Includes references listed for GenBank sequences. (DOCX 79 kb) Additional file 2: Table S2. Blood-feeding data for UK Culicoides exposed to membrane types. Number of Culicoides of each physiological state recorded following exposure to blood-feeding apparatus. (XLSX 21 kb) Abbreviations AIC: Akaike information criterion; BIC: Bayesian information criterion; BIN: Barcode index number; BTV: Bluetongue virus; DNA: Deoxyribonucleic acid; HKY+G: Hasegawa-Kishino-Yano with gamma-distribution rates; cox1: Mitochondrial cytochrome c oxidase subunit 1 gene; OVI: Onderstepoort Veterinary Institute; RDF: Roehl haplotype data files; RH: Relative humidity; RNA: Ribonucleic acid; RT-qPCR: Quantitative reverse transcription polymerase chain reaction; SBV: Schmallenberg virus
12 Barber et al. Parasites & Vectors (2018) 11:116 Page 12 of 13 Acknowledgements We acknowledge Nonito Pages, Maria Goffredo, Thomas Balenghien and Claire Garros for useful discussions on the subject of the manuscript. We also acknowledge provisionofthesbvstrainusedinexperimentsbyizsteramo(federicamonaco) from an isolate originally made available by the Institute of Diagnostic Virology, Friedrich-Loeffler-Institut, Riems, Germany (Martin Beer/Bernd Hoffmann). Funding This study was funded by EU grant FP EDENext ( SC was additionally funded by BBSRC grant BBS/E/I/ Availability of data and materials All data generated or analysed during this study are included in this published article and its additional files, with the exception of barcode sequence data which are listed by GenBank accession number. DNA sequences including electropherograms have been made publically available via the Barcode of Life Data System (BOLD) as data set DS-CUSBV (dx.doi.org/ /ds-cusbv). DNA sequences are also available in the GenBank database under accession numbers KT KT Authors contributions JB performed studies, edited and approved submission. LH carried out phylogenetic analysis, wrote and approved submission. RS processed samples, edited and approved submission. EV supervised studies, contributed to experimental design and edited and approved submission. SG carried out statistical analyses and edited and approved submission. KB carried out DNA sequencing and edited and approved submission. SC produced the experimental design, supervised the study and wrote and approved the submission. All authors read and approved the final manuscript. Ethics approval and consent to participate Chicks used in the trials were purchased dead and frozen from a local pet shop and blood was sourced from a commercial supplier (TCS Biosciences). No technique used during the trial required ethical approval. Consent for publication Not applicable. Competing interests The authors declare that they have no competing interests. Publisher s Note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. Author details 1 Vector-borne Viral Disease Programme, The Pirbright Institute, Pirbright, Surrey, UK. 2 National Centre for Vector Entomology, Institute of Parasitology, University of Zürich, Winterthurerstr. 266a, 8057 Zürich, Switzerland. Received: 22 April 2017 Accepted: 16 January 2018 References 1. Hoffmann B, Scheuch M, Höper D, Jungblut R, Holsteg M, Schirrmeier H, et al. Novel Orthobunyavirus in cattle, Europe Emerg Infect Dis. 2012;18: Doceul V, Lara E, Sailleau C, Belbis G, Richardson J, Breard E, et al. Epidemiology, molecular virology and diagnostics of Schmallenberg virus, an emerging Orthobunyavirus in Europe. Vet Res. 2013;44: Purse BV, Carpenter S, Venter GJ, Bellis G, Mullens BA. Bionomics of temperate and tropical Culicoides midges: Knowledge gaps and consequences for transmission of Culicoides-borne viruses. Annu Rev Entomol. 2014;60: Carpenter S, Groschup MH, Garros C, Felippe-Bauer ML, Purse BV. Culicoides biting midges, arboviruses and public health in Europe. Antiviral Res. 2013; 100(1): King B, O'Shea Brown T, Tarlinton R, Daly JM. Seroprevalence of Schmallenberg virus in the United Kingdom and the Republic of Ireland: Vet Microbiol. 2015;180(1 2): Barrett D, More SJ, O'Neill R, Bradshaw B, Casey M, Keane M, et al. Prevalence and distribution of exposure to Schmallenberg virus in Irish cattle during October 2012 to November BMC Vet Res. 2015; 11(1): Chenais E, Stahl K, Frossling J, Blomqvist G, Naslund K, Svensson L, et al. Schmallenberg virus beyond latitude 65 degrees N. Transbound Emerg Dis. 2015;62(5):e Meroc E, Poskin A, Van Loo H, Van Driessche E, Czaplicki G, Quinet C, et al. Follow-up of the Schmallenberg virus seroprevalence in Belgian cattle. Transbound Emerg Dis. 2015;62(5):e Wernike K, Holsteg M, Sasserath M, Beer M. Schmallenberg virus antibody development and decline in a naturally infected dairy cattle herd in Germany, Vet Microbiol. 2015;181(3 4): Surveillance SBV Congenital - Saison 2015/2016 Bilan au 18 julliet 2016 [ Surveillance%20SBV%20congénital_ _Traitement%204.pdf] Accessed on 19/12/ VMD authorises SBV vaccine for use in the UK. Vet Record. 2013;172(21): Carpenter S, Wilson A, Mellor PS. Culicoides and the emergence of bluetongue virus in northern Europe. Trends Microbiol. 2009;17(4): Yanase T, Kato T, Aizawa M, Shuto Y, Shirafuji H, Yamakawa M, Tsuda T. Genetic reassortment between Sathuperi and Shamonda viruses of the genus Orthobunyavirus in nature: implications for their genetic relationship to Schmallenberg virus. Arch Virol. 2012;157(8): Balenghien T, Pages N, Goffredo M, Carpenter S, Augot D, Jacquier E, et al. The emergence of Schmallenberg virus across Culicoides communities and ecosystems in Europe. Prev Vet Med. 2014;116(4): Elbers AR, Meiswinkel R, van Weezep E, van Oldruitenborgh-Oosterbaan MM, Kooi EA. Schmallenberg virus in Culicoides spp. biting midges, the Netherlands, Emerg Infect Dis. 2013;19(1): Elbers AR, Meiswinkel R, van Weezep E, Kooi EA, van der Poel WH. Schmallenberg virus in Culicoides biting midges in the Netherlands in Transbound Emerg Dis. 2015;62(3): De Regge N, Deblauwe I, De Deken R, Vantieghem P, Madder M, Geysen D, et al. Detection of Schmallenberg virus in different Culicoides spp. by realtime RT-PCR. Transbound Emerg Dis. 2012;59(6): De Regge N, De Deken R, Fassotte C, Losson B, Deblauwe I, Madder M, et al. Culicoides monitoring in Belgium in 2011: analysis of spatiotemporal abundance, species diversity and Schmallenberg virus detection. Med Veterinary Entomol. 2015;29(3): Wernike K, Jost H, Becker N, Schmidt-Chanasit J, Beer M. Lack of evidence for the presence of Schmallenberg virus in mosquitoes in Germany, Parasit Vectors. 2014;7: Veronesi E, Henstock M, Gubbins S, Batten C, Manley R, Barber J, et al. Implicating Culicoides biting midges as vectors of Schmallenberg virus using semi-quantitative RT-PCR. PLoS One. 2013;8(3):e Manley R, Harrup LE, Veronesi E, Stubbins F, Stoner J, Gubbins S, et al. Testing of UK populations of Culex pipiens L. for Schmallenberg virus vector competence and their colonization. PLoS One. 2015;10(8):e Meiswinkel R, Scolamacchia F, Dik M, Mudde J, Dijkstra E, Van Der Ven IJ, Elbers AR. The Mondrian matrix: Culicoides biting midge abundance and seasonal incidence during the epidemic of bluetongue in the Netherlands. Med Vet Entomol. 2014;28(1): Searle KR, Barber J, Stubbins F, Labuschagne K, Carpenter S, Butler A, et al. Environmental drivers of Culicoides phenology: How important is speciesspecific variation when determining disease policy? PLoS One. 2014;9(11): e Campbell JA, Pelham-Clinton EC. Taxonomic review of the British species of Culicoides Latreille (Diptera, Ceratopogonidae). Proc Roy Entomol Soc Lon (B). 1960;67: Nolan DV, Carpenter S, Barber J, Mellor PS, Dallas JF, Mordue Luntz AJ, Piertney SB. Rapid diagnostic PCR assays for members of the Culicoides obsoletus and Culicoides pulicaris species complexes, implicated vectors of bluetongue virus in Europe. Vet Microbiol. 2007;124(1 2): Schwenkenbecher JM, Mordue AJ, Piertney SB. Phylogenetic analysis indicates that Culicoides dewulfi should not be considered part of the Culicoides obsoletus complex. Bull Entomol Res. 2009;99(4): Kirkeby C, Dominiak P. Culicoides (Avaritia) gornostaevae Mirzaeva, 1984 (Diptera: Ceratopogonidae) - a possible vector species of the Obsoletus group new to the European fauna. Parasit Vectors. 2014;7:445.
13 Barber et al. Parasites & Vectors (2018) 11:116 Page 13 of Meiswinkel R, De Bree F, Bossers-De Vries R, Elbers AR. An unrecognized species of the Culicoides obsoletus complex feeding on livestock in the Netherlands. Vet Parasitol. 2015;207(3 4, 324): Pettersson E, Bensch S, Ander M, Chirico J, Sigvald R, Ignell R. Molecular identification of bloodmeals and species composition in Culicoides biting midges. Med Vet Entomol. 2013;27(1): Carpenter S, Lunt HL, Arav D, Venter GJ, Mellor PS. Oral susceptibility to bluetongue virus of Culicoides (Diptera: Ceratopogonidae) from the United Kingdom. J Med Entomol. 2006;43(1): Venter GJ, Paweska JT, Lunt H, Mellor PS, Carpenter S. An alternative method of blood-feeding Culicoides imicola and other haematophagous Culicoides species for vector competence studies. Vet Parasitol. 2005; 131(3 4): Nayduch D, Cohnstaedt LW, Saski C, Lawson D, Kersey P, Fife M, Carpenter S. Studying Culicoides vectors of BTV in the post-genomic era: Resources, bottlenecks to progress and future directions. Virus Res. 2014;182: Venter GJ, Paweska JT, van Dijk AA, Mellor PS, Tabachnick WJ. Vector competence of Culicoides bolitinos and C. imicola for South African bluetongue virus serotypes 1, 3 and 4. Med Vet Entomol. 1998;12(4): Dyce AL. The recognition of nulliparous and parous Culicoides (Diptera: Ceratopogonidae) without dissection. J Aust Entomol Soc. 1969;8: Harrup LE, Purse BP, Golding N, Mellor PS, Carpenter S. Larval development and emergence sites of farm-associated Culicoides (Diptera: Ceratopogonidae) in the United Kingdom. Med Vet Entomol. 2013;27(4): Hebert PD, Cywinska A, Ball SL, dewaard JR. Biological identifications through DNA barcodes. Proc Biol Sci. 2003;270(1512): Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotech. 1994;3: Hebert PDN, Penton EH, Burns JM, Janzen DH, Hallwach W. Ten species in one: DNA barcoding reveals cryptic species in the Neotropical skipper butterfly Astraptes fulgerator. Proc Nat Acad Sci USA. 2004;101: Ratnasingham S, Hebert PDN. BOLD: The Barcode of Life Data System. Mol Ecol Notes. 2007;7: Madden T. The BLAST Sequence Analysis Tool, avaliable at nih.gov/books/nbk In: The NCBI Handbook. Edited by McEntyre J, Ostell J. Bethesda MD, USA: National Center for Biotechnology Information; Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32(5): Penn O, Privman E, Landan G, Graur D, Pupko T. An alignment confidence score capturing robustness to guide-tree uncertainty. Mol Biol Evol. 2010; 27(8): Darriba D, Taboada GL, Doallo R, Posada D. jmodeltest 2: more models, new heuristics and parallel computing. Nat Methods. 2012;9(8): Guindon S, Gascuel O. A simple, fast and accurate method to estimate large phylogenies by maximum-likelihood. Systematic Biol. 2003;52: Huelsenbeck JP, Ronquist F. MRBAYES: Bayesian inference of phylogeny. Bioinformatics. 2001;17: Ronquist F, Huelsenbeck JP. MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003;19: Hasegawa M, Kishino H, Yano T. Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol. 1985;22(2): Harrup LE, Laban S, Purse BV, Reddy YK, Reddy YN, Byregowda SM, et al. DNA barcoding and surveillance sampling strategies for Culicoides biting midges (Diptera: Ceratopogonidae) in southern India. Parasit Vectors. 2016;9: Ratnasingham S, Hebert PDN. A DNA-based registry for all animal species: The Barcode Index Number (BIN) system. PLoS One. 2013;8(7):e Nylander JAA, Wilgenbusch JC, Warren DL, Swofford DL. AWTY: A system for graphical exploration of MCMC convergence in Bayesian phylogenetics. Bioinformatics. 2008;24(4): Paradis E, Claude J, Strimmer K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics. 2004;20: Revell LJ. phytools: An R package for phylogenetic comparative biology (and other things). Meth Ecol Evol. 2012;3: R Development Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. ISBN , edn Librado P, Rozas J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics. 2009;25: Fluxus-Engineering. Network version , avaliable at: Bandelt H-J, Forster P, Röhl A. Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol. 1999;16: Polzin T, Daneschmand SV. On Steiner trees and minimum spanning trees in hypergraphs. Oper Res Lett. 2003;31: Brown SD, Collins RA, Boyer S, Lefort MC, Malumbres-Olarte J, Vink CJ, Cruickshank RH. Spider: an R package for the analysis of species identity and evolution, with particular reference to DNA barcoding. Mol Ecol Resour. 2012;12(3): Wenk CE, Kaufmann C, Schaffner F, Mathis A. Molecular characterization of Swiss Ceratopogonidae (Diptera) and evaluation of real-time PCR assays for the identification of Culicoides biting midges. Vet Parasitol. 2012;184(2 4): Jennings DM, Mellor PS. The vector potential of British Culicoides species for bluetongue virus. Vet Microbiol. 1988;17(1): Blackwell A, Mellor PS, Mordue W. Laboratory feeding of Culicoides impunctatus (Diptera: Ceratopogonidae) through natural and artificial membranes. Med Vet Entomol. 1994;31(2): Blackwell A, Mellor PS, Mordue W. Methods for enhancing the blood feeding response of field-collected Culicoides impunctatus (Diptera: Ceratopogonidae). J Med Entomol. 1996;33(3): Gubbins S, Richardson J, Baylis M, Wilson AJ, Abrahantes JC. Modelling the continental-scale spread of Schmallenberg virus in Europe: approaches and challenges. Prev Vet Med. 2014;116(4): Gubbins S, Turner J, Baylis M, van der Stede Y, van Schaik G, Abrahantes JC, Wilson AJ. Inferences about the transmission of Schmallenberg virus within and between farms. Prev Vet Med. 2014;116(4): Ander M, Troell K, Chirico J. Barcoding of biting midges in the genus Culicoides: a tool for species determination. Med Vet Entomol. 2013;27(3): Mathieu B, Delécolle JC, Garros C, Balenghien T, Setier-Rio ML, Candolfi E, Cêtre-Sossah C. Simultaneous quantification of the relative abundance of species complex members: application to Culicoides obsoletus and Culicoides scoticus (Diptera: Ceratopogonidae), potential vectors of bluetongue virus. Vet Parasitol. 2011;182(2 4): Gomulski LM, Meiswinkel R, Delécolle JC, Goffredo M, Gasperi G. Phylogenetic relationships of the sub-genus Avaritia Fox, 1995 including Culicoides obsoletus (Diptera, Ceratopogonidae) in Italy based on internal transcribed spacer 2 ribosomal DNA sequences ;10: Harrup L, Bellis GA, Balenghien T, Garros C. Culicoides Latreille (Diptera: Ceratopogonidae) taxonomy: Current challenges and future directions. Infect Genet Evol. 2015;30: Linton YM, Mordue AJ, Cruickshank RH, Meiswinkel R, Mellor PS, Dallas JF. Phylogenetic analysis of the mitochondrial cytochrome oxidase subunit I gene of five species of the Culicoides imicola species complex. Med Vet Entomol. 2002;16: Submit your next manuscript to BioMed Central and we will help you at every step: We accept pre-submission inquiries Our selector tool helps you to find the most relevant journal We provide round the clock customer support Convenient online submission Thorough peer review Inclusion in PubMed and all major indexing services Maximum visibility for your research Submit your manuscript at
Systematics and taxonomy of the genus Culicoides what is coming next?
 Systematics and taxonomy of the genus Culicoides what is coming next? Claire Garros 1, Bruno Mathieu 2, Thomas Balenghien 1, Jean-Claude Delécolle 2 1 CIRAD, Montpellier, France 2 IPPTS, Strasbourg, France
Systematics and taxonomy of the genus Culicoides what is coming next? Claire Garros 1, Bruno Mathieu 2, Thomas Balenghien 1, Jean-Claude Delécolle 2 1 CIRAD, Montpellier, France 2 IPPTS, Strasbourg, France
Transmission of the virus (SBV) Stéphan Zientara UMR 1161 ANSES/INRA/ENVA
 Transmission of the virus (SBV) Stéphan Zientara UMR 1161 ANSES/INRA/ENVA April 2, 2012 Transmission routes Direct transmission Vertical transmission Insect transmission Detection of Schmallenberg virus
Transmission of the virus (SBV) Stéphan Zientara UMR 1161 ANSES/INRA/ENVA April 2, 2012 Transmission routes Direct transmission Vertical transmission Insect transmission Detection of Schmallenberg virus
Culicoides species composition and abundance on Irish cattle farms: implications for arboviral disease transmission
 Collins et al. Parasites & Vectors (2018) 11:472 https://doi.org/10.1186/s13071-018-3010-6 RESEARCH Culicoides species composition and abundance on Irish cattle farms: implications for arboviral disease
Collins et al. Parasites & Vectors (2018) 11:472 https://doi.org/10.1186/s13071-018-3010-6 RESEARCH Culicoides species composition and abundance on Irish cattle farms: implications for arboviral disease
Danish Culicoides species of the Obsoletus group identified by morphological methods
 Danish Culicoides species of the Obsoletus group identified by morphological methods Søren Achim Nielsen Dept of Environmental, Social and Spatial Change Roskilde University Denmark Michael Kristensen
Danish Culicoides species of the Obsoletus group identified by morphological methods Søren Achim Nielsen Dept of Environmental, Social and Spatial Change Roskilde University Denmark Michael Kristensen
Culicoides and the global epidemiology of bluetongue virus infection
 Vet. Ital., 40 (3), 145-150 Epidemiology and vectors Culicoides and the global epidemiology of bluetongue virus infection W.J. Tabachnick Florida Medical Entomology Laboratory, Department of Entomology
Vet. Ital., 40 (3), 145-150 Epidemiology and vectors Culicoides and the global epidemiology of bluetongue virus infection W.J. Tabachnick Florida Medical Entomology Laboratory, Department of Entomology
A comparison of commercial light-emitting diode baited suction traps for surveillance of Culicoides in northern Europe
 Hope et al. Parasites & Vectors (2015) 8:239 DOI 10.1186/s13071-015-0846-x RESEARCH Open Access A comparison of commercial light-emitting diode baited suction traps for surveillance of Culicoides in northern
Hope et al. Parasites & Vectors (2015) 8:239 DOI 10.1186/s13071-015-0846-x RESEARCH Open Access A comparison of commercial light-emitting diode baited suction traps for surveillance of Culicoides in northern
Implicating Culicoides Biting Midges as Vectors of Schmallenberg Virus Using Semi-Quantitative RT-PCR
 Implicating Culicoides Biting Midges as Vectors of Schmallenberg Virus Using Semi-Quantitative RT-PCR Eva Veronesi 1, Mark Henstock 1, Simon Gubbins 1, Carrie Batten 1, Robyn Manley 1, James Barber 1,
Implicating Culicoides Biting Midges as Vectors of Schmallenberg Virus Using Semi-Quantitative RT-PCR Eva Veronesi 1, Mark Henstock 1, Simon Gubbins 1, Carrie Batten 1, Robyn Manley 1, James Barber 1,
Environmental Drivers of Culicoides Phenology: How Important Is Species-Specific Variation When Determining Disease Policy?
 Environmental Drivers of Culicoides Phenology: How Important Is Species-Specific Variation When Determining Disease Policy? Kate R. Searle 1 *, James Barber 2, Francesca Stubbins 2, Karien Labuschagne
Environmental Drivers of Culicoides Phenology: How Important Is Species-Specific Variation When Determining Disease Policy? Kate R. Searle 1 *, James Barber 2, Francesca Stubbins 2, Karien Labuschagne
G. Kluiters 1*, N. Pagès 2,7, S. Carpenter 3, L. Gardès 4,5, H. Guis 4,5, M. Baylis 1,6 and C. Garros 4,5
 Kluiters et al. Parasites & Vectors (2016) 9:262 DOI 10.1186/s13071-016-1520-7 RESEARCH Open Access Morphometric discrimination of two sympatric sibling species in the Palaearctic region, Culicoides obsoletus
Kluiters et al. Parasites & Vectors (2016) 9:262 DOI 10.1186/s13071-016-1520-7 RESEARCH Open Access Morphometric discrimination of two sympatric sibling species in the Palaearctic region, Culicoides obsoletus
Sheep breed and shearing influences attraction and blood-feeding behaviour of Culicoides (Diptera: Ceratopogonidae) on a UK farm
 Hope et al. Parasites & Vectors (2018) 11:473 https://doi.org/10.1186/s13071-018-3003-5 RESEARCH Open Access Sheep breed and shearing influences attraction and blood-feeding behaviour of Culicoides (Diptera:
Hope et al. Parasites & Vectors (2018) 11:473 https://doi.org/10.1186/s13071-018-3003-5 RESEARCH Open Access Sheep breed and shearing influences attraction and blood-feeding behaviour of Culicoides (Diptera:
The influence of temperature and humidity on the flight activity of Culicoides imicola both under laboratory and field conditions
 Venter et al. Parasites & Vectors (2019) 12:4 https://doi.org/10.1186/s13071-018-3272-z RESEARCH The influence of temperature and humidity on the flight activity of Culicoides imicola both under laboratory
Venter et al. Parasites & Vectors (2019) 12:4 https://doi.org/10.1186/s13071-018-3272-z RESEARCH The influence of temperature and humidity on the flight activity of Culicoides imicola both under laboratory
Progress and knowledge gaps in Culicoides genetics, genomics and population modelling: 2003 to 2014
 Progress and knowledge gaps in Culicoides genetics, genomics and population modelling: 2003 to 2014 Simon Carpenter Vector borne Disease Programme, The Pirbright Institute, United Kingdom Corresponding
Progress and knowledge gaps in Culicoides genetics, genomics and population modelling: 2003 to 2014 Simon Carpenter Vector borne Disease Programme, The Pirbright Institute, United Kingdom Corresponding
* * *Determine Culicoides spp. present in the Southeast, including at
 Stacey Vigil, Joseph L. Corn, Mark G. Ruder, and David K. Stallknecht svigil@uga.edu Southeast Cooperative Wildlife Disease Study, College of Veterinary Medicine, University of Georgia United States Animal
Stacey Vigil, Joseph L. Corn, Mark G. Ruder, and David K. Stallknecht svigil@uga.edu Southeast Cooperative Wildlife Disease Study, College of Veterinary Medicine, University of Georgia United States Animal
Characterizing the species composition of European Culicoides vectors by means of the Köppen-Geiger climate classification
 Brugger and Rubel Parasites & Vectors 2013, 6:333 SHORT REPORT Open Access Characterizing the species composition of European Culicoides vectors by means of the Köppen-Geiger climate classification Katharina
Brugger and Rubel Parasites & Vectors 2013, 6:333 SHORT REPORT Open Access Characterizing the species composition of European Culicoides vectors by means of the Köppen-Geiger climate classification Katharina
EXTERNAL SCIENTIFIC REPORT
 EXTERNAL SCIENTIFIC REPORT APPROVED: 8 February 2017 doi:10.2903/sp.efsa.2017.en-1182 A first estimation of Culicoides imicola and Culicoides obsoletus/culicoides scoticus seasonality and abundance in
EXTERNAL SCIENTIFIC REPORT APPROVED: 8 February 2017 doi:10.2903/sp.efsa.2017.en-1182 A first estimation of Culicoides imicola and Culicoides obsoletus/culicoides scoticus seasonality and abundance in
DNA barcoding and surveillance sampling strategies for Culicoides biting midges (Diptera: Ceratopogonidae) in southern India
 Harrup et al. Parasites & Vectors (2016) 9:461 DOI 10.1186/s13071-016-1722-z RESEARCH DNA barcoding and surveillance sampling strategies for Culicoides biting midges (Diptera: Ceratopogonidae) in southern
Harrup et al. Parasites & Vectors (2016) 9:461 DOI 10.1186/s13071-016-1722-z RESEARCH DNA barcoding and surveillance sampling strategies for Culicoides biting midges (Diptera: Ceratopogonidae) in southern
Schmallenberg Virus Infections in Ruminants
 Schmallenberg Virus Infections in Ruminants F. J. Conraths, B. Hoffmann, D. Höper, M. Scheuch, R. Jungblut, M. Holsteg, H. Schirrmeier, M. Eschbaumer, K. Goller, K. Wernike, M. Fischer, A. Breithaupt,
Schmallenberg Virus Infections in Ruminants F. J. Conraths, B. Hoffmann, D. Höper, M. Scheuch, R. Jungblut, M. Holsteg, H. Schirrmeier, M. Eschbaumer, K. Goller, K. Wernike, M. Fischer, A. Breithaupt,
WAGENINGEN UNIVERSITY LABORATORY OF ENTOMOLOGY
 WAGENINGEN UNIVERSITY LABORATORY OF ENTOMOLOGY The overwintering behaviour of adult Culicoides species on livestock farms in the Netherlands and the effect of indoor insecticidal treatment on Culicoides
WAGENINGEN UNIVERSITY LABORATORY OF ENTOMOLOGY The overwintering behaviour of adult Culicoides species on livestock farms in the Netherlands and the effect of indoor insecticidal treatment on Culicoides
Culicoides midges (Diptera: Ceratopogonidae) as vectors of orbiviruses in Slovakia
 Culicoides midges (Diptera: Ceratopogonidae) as vectors of orbiviruses in Slovakia Adela Sarvašová 1, Maria Goffredo 2, Igor Sopoliga 3, Giovanni Savini 2 & Alica Kočišová 1* 1 University of Veterinary
Culicoides midges (Diptera: Ceratopogonidae) as vectors of orbiviruses in Slovakia Adela Sarvašová 1, Maria Goffredo 2, Igor Sopoliga 3, Giovanni Savini 2 & Alica Kočišová 1* 1 University of Veterinary
Lecture 11 Wednesday, September 19, 2012
 Lecture 11 Wednesday, September 19, 2012 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean
Lecture 11 Wednesday, September 19, 2012 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean
Veterinary Diagnostics Portfolio Overview. Complete solutions for veterinary testing and pathogen research
 Veterinary Diagnostics Portfolio Overview Complete solutions for veterinary testing and pathogen research Sample preparation products Cat. no. (number of preps) Target analyte Product Short description
Veterinary Diagnostics Portfolio Overview Complete solutions for veterinary testing and pathogen research Sample preparation products Cat. no. (number of preps) Target analyte Product Short description
Möhlmann et al. Parasites & Vectors (2018) 11:217
 Möhlmann et al. Parasites & Vectors (2018) 11:217 https://doi.org/10.1186/s13071-018-2792-x RESEARCH Open Access Community analysis of the abundance and diversity of biting midge species (Diptera: Ceratopogonidae)
Möhlmann et al. Parasites & Vectors (2018) 11:217 https://doi.org/10.1186/s13071-018-2792-x RESEARCH Open Access Community analysis of the abundance and diversity of biting midge species (Diptera: Ceratopogonidae)
Seroprevalence of antibodies to Schmallenberg virus in livestock
 Seroprevalence of antibodies to Schmallenberg virus in livestock Armin R.W. Elbers Dept. Epidemiology, Crisis organisation and Diagnostics Central Veterinary Institute (CVI) part of Wageningen UR armin.elbers@wur.nl
Seroprevalence of antibodies to Schmallenberg virus in livestock Armin R.W. Elbers Dept. Epidemiology, Crisis organisation and Diagnostics Central Veterinary Institute (CVI) part of Wageningen UR armin.elbers@wur.nl
Role of different Culicoides vectors (Diptera: Ceratopogonidae) in bluetongue virus transmission and overwintering in Sardinia (Italy)
 Foxi et al. Parasites & Vectors (2016) 9:440 DOI 10.1186/s13071-016-1733-9 RESEARCH Open Access Role of different Culicoides vectors (Diptera: Ceratopogonidae) in bluetongue virus transmission and overwintering
Foxi et al. Parasites & Vectors (2016) 9:440 DOI 10.1186/s13071-016-1733-9 RESEARCH Open Access Role of different Culicoides vectors (Diptera: Ceratopogonidae) in bluetongue virus transmission and overwintering
Veterinary Parasitology
 Veterinary Parasitology 184 (2012) 258 266 Contents lists available at SciVerse ScienceDirect Veterinary Parasitology jou rn al h om epa ge: www.elsevier.com/locate/vetpar Molecular characterization of
Veterinary Parasitology 184 (2012) 258 266 Contents lists available at SciVerse ScienceDirect Veterinary Parasitology jou rn al h om epa ge: www.elsevier.com/locate/vetpar Molecular characterization of
Investigation of Culicoides spp. preference for light colour and source using light emitting diodes and fluorescent light
 514 Investigation of Culicoides spp. preference for light colour and source using light emitting diodes and fluorescent light A.B. Jenkins and M.B. Young # Animal and Poultry Science, School of Agricultural
514 Investigation of Culicoides spp. preference for light colour and source using light emitting diodes and fluorescent light A.B. Jenkins and M.B. Young # Animal and Poultry Science, School of Agricultural
Indoor and outdoor winter activity of Culicoides biting midges, vectors of bluetongue virus, in Italy
 Medical and Veterinary Entomology (2018) 32, 70 77 doi: 10.1111/mve.12260 Indoor and outdoor winter activity of Culicoides biting midges, vectors of bluetongue virus, in Italy A. MAGLIANO 1, P. SCARAMOZZINO
Medical and Veterinary Entomology (2018) 32, 70 77 doi: 10.1111/mve.12260 Indoor and outdoor winter activity of Culicoides biting midges, vectors of bluetongue virus, in Italy A. MAGLIANO 1, P. SCARAMOZZINO
PCR detection of Leptospira in. stray cat and
 PCR detection of Leptospira in 1 Department of Pathology, School of Veterinary Medicine, Islamic Azad University, Shahrekord Branch, Shahrekord, Iran 2 Department of Microbiology, School of Veterinary
PCR detection of Leptospira in 1 Department of Pathology, School of Veterinary Medicine, Islamic Azad University, Shahrekord Branch, Shahrekord, Iran 2 Department of Microbiology, School of Veterinary
The Culicoides obsoletus group in Italy: relative abundance, geographic range, and role as vector for Bluetongue virus
 The Culicoides obsoletus group in Italy: relative abundance, geographic range, and role as vector for Bluetongue virus Maria Goffredo 1*, Rudy Meiswinkel, Valentina Federici 1, Francesca Di Nicola 1, Giuseppe
The Culicoides obsoletus group in Italy: relative abundance, geographic range, and role as vector for Bluetongue virus Maria Goffredo 1*, Rudy Meiswinkel, Valentina Federici 1, Francesca Di Nicola 1, Giuseppe
Testing Phylogenetic Hypotheses with Molecular Data 1
 Testing Phylogenetic Hypotheses with Molecular Data 1 How does an evolutionary biologist quantify the timing and pathways for diversification (speciation)? If we observe diversification today, the processes
Testing Phylogenetic Hypotheses with Molecular Data 1 How does an evolutionary biologist quantify the timing and pathways for diversification (speciation)? If we observe diversification today, the processes
Culicoides DISEASE TRANSMISSION. Arthropod vectors Culicoides
 Culicoides Author: Dr. Gert Venter Licensed under a Creative Commons Attribution license. DISEASE TRANSMISSION In 1943 Du Toit conducted the first successful transmission of BTV from infected Culicoides
Culicoides Author: Dr. Gert Venter Licensed under a Creative Commons Attribution license. DISEASE TRANSMISSION In 1943 Du Toit conducted the first successful transmission of BTV from infected Culicoides
Epidemiology and vectors Vet. Ital., 40 (3), & R. Meiswinkel
 Vet. Ital., 40 (3), 260-265 Entomological surveillance of bluetongue in Italy: methods of capture, catch analysis and identification of Culicoides biting midges M. Goffredo (1) (1, 2) & R. Meiswinkel (1)
Vet. Ital., 40 (3), 260-265 Entomological surveillance of bluetongue in Italy: methods of capture, catch analysis and identification of Culicoides biting midges M. Goffredo (1) (1, 2) & R. Meiswinkel (1)
Kirkeby, Carsten Thure; Dominiak, Patrycja. Published in: Parasites & Vectors. Link to article, DOI: / Publication date: 2014
 Downloaded from orbit.dtu.dk on: Jan 26, 2018 Culicoides (Avaritia) gornostaevae Mirzaeva, 1984 (Diptera: Ceratopogonidae) a possible vector species of the Obsoletus group new to the European fauna. Kirkeby,
Downloaded from orbit.dtu.dk on: Jan 26, 2018 Culicoides (Avaritia) gornostaevae Mirzaeva, 1984 (Diptera: Ceratopogonidae) a possible vector species of the Obsoletus group new to the European fauna. Kirkeby,
Culicoides species from the subgenus Culicoides in Catalonia (NE Spain)
 Culicoides species from the subgenus Culicoides in Catalonia (NE Spain) Pagès, N., Muñoz-Muñoz, F., Talavera, S., Sarto, V., Lorca, C. and Nuñez, J.I. Identification Background Identification of Culicoides
Culicoides species from the subgenus Culicoides in Catalonia (NE Spain) Pagès, N., Muñoz-Muñoz, F., Talavera, S., Sarto, V., Lorca, C. and Nuñez, J.I. Identification Background Identification of Culicoides
RISK ASSESSMENT WORKPACKAGE 5 BTV OVERWINTERING BY HORIZONTAL TRANSMISSION IN VECTORS, RUMINANTS OR IN BOTH
 WORKPACKAGE 5 RISK ASSESSMENT S. Napp A. Alba I. García A. Allepuz J. Casal BTV OVERWINTERING BY HORIZONTAL TRANSMISSION IN VECTORS, RUMINANTS OR IN BOTH P. Calistri A. Giovannini S. Gubbins INTRODUCTION
WORKPACKAGE 5 RISK ASSESSMENT S. Napp A. Alba I. García A. Allepuz J. Casal BTV OVERWINTERING BY HORIZONTAL TRANSMISSION IN VECTORS, RUMINANTS OR IN BOTH P. Calistri A. Giovannini S. Gubbins INTRODUCTION
OIE Collaborating Centre for Training in. Integrated Livestock and Wildlife Health and Management, Onderstepoort. Development of the Centre
 OIE Collaborating Centre for Training in Integrated Livestock and Wildlife Health and Management, Onderstepoort Development of the Centre Consortium Partner Institutions Proposal - OIE Collaboration Centre
OIE Collaborating Centre for Training in Integrated Livestock and Wildlife Health and Management, Onderstepoort Development of the Centre Consortium Partner Institutions Proposal - OIE Collaboration Centre
Epidemiological analysis of the 2006 bluetongue virus serotype 8 epidemic in north-western Europe. Within herd distribution of infection
 Epidemiological analysis of the 26 bluetongue virus serotype 8 epidemic in north-western Europe Within herd distribution of infection A.R.W. Elbers 1, K. Mintiens 2, G. Gerbier 3, A.N. van der Spek 4,
Epidemiological analysis of the 26 bluetongue virus serotype 8 epidemic in north-western Europe Within herd distribution of infection A.R.W. Elbers 1, K. Mintiens 2, G. Gerbier 3, A.N. van der Spek 4,
3. records of distribution for proteins and feeds are being kept to facilitate tracing throughout the animal feed and animal production chain.
 CANADA S FEED BAN The purpose of this paper is to explain the history and operation of Canada s feed ban and to put it into a broader North American context. Canada and the United States share the same
CANADA S FEED BAN The purpose of this paper is to explain the history and operation of Canada s feed ban and to put it into a broader North American context. Canada and the United States share the same
GEODIS 2.0 DOCUMENTATION
 GEODIS.0 DOCUMENTATION 1999-000 David Posada and Alan Templeton Contact: David Posada, Department of Zoology, 574 WIDB, Provo, UT 8460-555, USA Fax: (801) 78 74 e-mail: dp47@email.byu.edu 1. INTRODUCTION
GEODIS.0 DOCUMENTATION 1999-000 David Posada and Alan Templeton Contact: David Posada, Department of Zoology, 574 WIDB, Provo, UT 8460-555, USA Fax: (801) 78 74 e-mail: dp47@email.byu.edu 1. INTRODUCTION
Phylogeny Reconstruction
 Phylogeny Reconstruction Trees, Methods and Characters Reading: Gregory, 2008. Understanding Evolutionary Trees (Polly, 2006) Lab tomorrow Meet in Geology GY522 Bring computers if you have them (they will
Phylogeny Reconstruction Trees, Methods and Characters Reading: Gregory, 2008. Understanding Evolutionary Trees (Polly, 2006) Lab tomorrow Meet in Geology GY522 Bring computers if you have them (they will
Applied-for scope of designation and notification of a Conformity Assessment Body Regulation (EU) 2017/746 (IVDR)
 Ref. Ares(2018)2576484-17/05/2018 NBOG s Best Practice Guide applicable for MDR IVDR NBOG F 2017-4 This document has been endorsed by the Medical Device Coordination Group (MDCG) established by Article
Ref. Ares(2018)2576484-17/05/2018 NBOG s Best Practice Guide applicable for MDR IVDR NBOG F 2017-4 This document has been endorsed by the Medical Device Coordination Group (MDCG) established by Article
J. Med. Entomol. 44(6): 1019Ð1025 (2007)
 VECTOR CONTROL, PEST MANAGEMENT, RESISTANCE, REPELLENTS Molecular Identification of Western European Species of Obsoletus Complex (Diptera: Ceratopogonidae) by an Internal Transcribed Spacer-1 rdna Multiplex
VECTOR CONTROL, PEST MANAGEMENT, RESISTANCE, REPELLENTS Molecular Identification of Western European Species of Obsoletus Complex (Diptera: Ceratopogonidae) by an Internal Transcribed Spacer-1 rdna Multiplex
CLADISTICS Student Packet SUMMARY Phylogeny Phylogenetic trees/cladograms
 CLADISTICS Student Packet SUMMARY PHYLOGENETIC TREES AND CLADOGRAMS ARE MODELS OF EVOLUTIONARY HISTORY THAT CAN BE TESTED Phylogeny is the history of descent of organisms from their common ancestor. Phylogenetic
CLADISTICS Student Packet SUMMARY PHYLOGENETIC TREES AND CLADOGRAMS ARE MODELS OF EVOLUTIONARY HISTORY THAT CAN BE TESTED Phylogeny is the history of descent of organisms from their common ancestor. Phylogenetic
Genotypes of Cornel Dorset and Dorset Crosses Compared with Romneys for Melatonin Receptor 1a
 Genotypes of Cornell Dorset and Dorset Crosses Compared with Romneys for Melatonin Receptor 1a By Christian Posbergh Cornell Undergraduate Honor Student, Dept. Animal Science Abstract: Sheep are known
Genotypes of Cornell Dorset and Dorset Crosses Compared with Romneys for Melatonin Receptor 1a By Christian Posbergh Cornell Undergraduate Honor Student, Dept. Animal Science Abstract: Sheep are known
Vector-Borne Diseases, Surveillance, Prevention
 Vector-Borne Diseases, Surveillance, Prevention Journal of Medical Entomology, 53(2), 2016, 416 424 doi: 10.1093/jme/tjv197 Advance Access Publication Date: 22 December 2015 Research article Seasonal Dynamics,
Vector-Borne Diseases, Surveillance, Prevention Journal of Medical Entomology, 53(2), 2016, 416 424 doi: 10.1093/jme/tjv197 Advance Access Publication Date: 22 December 2015 Research article Seasonal Dynamics,
Bi156 Lecture 1/13/12. Dog Genetics
 Bi156 Lecture 1/13/12 Dog Genetics The radiation of the family Canidae occurred about 100 million years ago. Dogs are most closely related to wolves, from which they diverged through domestication about
Bi156 Lecture 1/13/12 Dog Genetics The radiation of the family Canidae occurred about 100 million years ago. Dogs are most closely related to wolves, from which they diverged through domestication about
Terrestrial and Aquatic Manuals and the mechanism of standard adoption
 Dr Patrick Bastiaensen Programme Officer OIE Sub-Regional Representation for Eastern Africa Terrestrial and Aquatic Manuals and the mechanism of standard adoption Presented during the Regional Workshop
Dr Patrick Bastiaensen Programme Officer OIE Sub-Regional Representation for Eastern Africa Terrestrial and Aquatic Manuals and the mechanism of standard adoption Presented during the Regional Workshop
Identification of field-caught Culicoides biting midges using matrix-assisted laser desorption/ionization time of flight mass spectrometry
 Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2012 Identification of field-caught Culicoides biting midges using matrix-assisted
Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2012 Identification of field-caught Culicoides biting midges using matrix-assisted
Christian Kaufmann *, Irene C Steinmann, Daniel Hegglin, Francis Schaffner and Alexander Mathis
 Kaufmann et al. Parasites & Vectors 22, 5:246 RESEARCH Open Access Spatio-temporal occurrence of Culicoides biting midges in the climatic regions of Switzerland, along with large scale species identification
Kaufmann et al. Parasites & Vectors 22, 5:246 RESEARCH Open Access Spatio-temporal occurrence of Culicoides biting midges in the climatic regions of Switzerland, along with large scale species identification
EUROPEAN COMMISSION HEALTH & CONSUMERS DIRECTORATE-GENERAL. Unit G5 - Veterinary Programmes
 EUROPEAN COMMISSION HEALTH & CONSUMERS DIRECTORATE-GENERAL Unit G5 - Veterinary Programmes SANCO/10853/2012 Programmes for the eradication, control and monitoring of certain animal diseases and zoonoses
EUROPEAN COMMISSION HEALTH & CONSUMERS DIRECTORATE-GENERAL Unit G5 - Veterinary Programmes SANCO/10853/2012 Programmes for the eradication, control and monitoring of certain animal diseases and zoonoses
COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST
 Big Idea 1 Evolution INVESTIGATION 3 COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST How can bioinformatics be used as a tool to determine evolutionary relationships and to
Big Idea 1 Evolution INVESTIGATION 3 COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST How can bioinformatics be used as a tool to determine evolutionary relationships and to
Species: Panthera pardus Genus: Panthera Family: Felidae Order: Carnivora Class: Mammalia Phylum: Chordata
 CHAPTER 6: PHYLOGENY AND THE TREE OF LIFE AP Biology 3 PHYLOGENY AND SYSTEMATICS Phylogeny - evolutionary history of a species or group of related species Systematics - analytical approach to understanding
CHAPTER 6: PHYLOGENY AND THE TREE OF LIFE AP Biology 3 PHYLOGENY AND SYSTEMATICS Phylogeny - evolutionary history of a species or group of related species Systematics - analytical approach to understanding
ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS. Medicinal product no longer authorised
 ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS 1 1. NAME OF THE VETERINARY MEDICINAL PRODUCT BTVPUR AlSap 1 suspension for injection for sheep and cattle. 2. QUALITATIVE AND QUANTITATIVE COMPOSITION Each dose
ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS 1 1. NAME OF THE VETERINARY MEDICINAL PRODUCT BTVPUR AlSap 1 suspension for injection for sheep and cattle. 2. QUALITATIVE AND QUANTITATIVE COMPOSITION Each dose
SCWDS HD Surveillance 11/8/2016. Update on SCWDS Culicoides Surveys in the Southeast. Common Culicoides species in the Southeast U.S.
 /8/0 Update on SCWDS Culicoides Surveys in the Southeast >00 sites >7,500 trap-nights WMAs, parks, etc July September CDC light traps Stacey Vigil, Mark Ruder, and Joseph L. Corn Southeastern Cooperative
/8/0 Update on SCWDS Culicoides Surveys in the Southeast >00 sites >7,500 trap-nights WMAs, parks, etc July September CDC light traps Stacey Vigil, Mark Ruder, and Joseph L. Corn Southeastern Cooperative
Feeding behaviour of Culicoides spp. (Diptera: Ceratopogonidae) on cattle and sheep in northeast Germany
 Ayllón et al. Parasites & Vectors 2014, 7:34 RESEARCH Open Access Feeding behaviour of Culicoides spp. (Diptera: Ceratopogonidae) on cattle and sheep in northeast Germany Tania Ayllón 1, Ard M Nijhof 1,
Ayllón et al. Parasites & Vectors 2014, 7:34 RESEARCH Open Access Feeding behaviour of Culicoides spp. (Diptera: Ceratopogonidae) on cattle and sheep in northeast Germany Tania Ayllón 1, Ard M Nijhof 1,
Entomological surveillance of bluetongue in France in 2002
 Vet. Ital., (3), 226-23 Entomological surveillance of bluetongue in France in 22 T. Baldet (), J.-C. Delécolle (2), B. Mathieu (3), S. de La Rocque () & F. Roger () () CIRAD-EMVT, TA 3 E, Campus International
Vet. Ital., (3), 226-23 Entomological surveillance of bluetongue in France in 22 T. Baldet (), J.-C. Delécolle (2), B. Mathieu (3), S. de La Rocque () & F. Roger () () CIRAD-EMVT, TA 3 E, Campus International
Identity and diversity of blood meal hosts of biting midges (Diptera: Ceratopogonidae: Culicoides Latreille) in Denmark
 Lassen et al. Parasites & Vectors 2012, 5:143 RESEARCH Identity and diversity of blood meal hosts of biting midges (Diptera: Ceratopogonidae: Culicoides Latreille) in Denmark Sandra B Lassen 1, Søren Achim
Lassen et al. Parasites & Vectors 2012, 5:143 RESEARCH Identity and diversity of blood meal hosts of biting midges (Diptera: Ceratopogonidae: Culicoides Latreille) in Denmark Sandra B Lassen 1, Søren Achim
Introduction to Biorisk and the OIE Standard
 Introduction to Biorisk and the OIE Standard World Association of Veterinary Laboratory Diagnosticians 18 th International Symposium, Sorrento, Italy 7 th -10 th June 2017 2015 Dr. Anthony Fooks Member,
Introduction to Biorisk and the OIE Standard World Association of Veterinary Laboratory Diagnosticians 18 th International Symposium, Sorrento, Italy 7 th -10 th June 2017 2015 Dr. Anthony Fooks Member,
Regional research activities and state of the art of Vmerge Project: Emerging viralvector
 Regional research activities and state of the art of Vmerge Project: Emerging viralvector borne diseases Joint permanent committee 4th November 2014 Cirad Key features of Vmerge Cirad - F Borne Objectives
Regional research activities and state of the art of Vmerge Project: Emerging viralvector borne diseases Joint permanent committee 4th November 2014 Cirad Key features of Vmerge Cirad - F Borne Objectives
Jean-Yves Zimmer a *, Bertrand Losson b, Claude Saegerman c, Eric Haubruge a & Frédéric Francis a
 Annales de la Société entomologique de France (N.S.), 2013 Vol. 49, No. 3, 335 344, http://dx.doi.org/10.1080/00379271.2013.854100 Breeding sites and species association of the main Bluetongue and Schmallenberg
Annales de la Société entomologique de France (N.S.), 2013 Vol. 49, No. 3, 335 344, http://dx.doi.org/10.1080/00379271.2013.854100 Breeding sites and species association of the main Bluetongue and Schmallenberg
Ticks and tick-borne pathogens Jordi Tarrés-Call, Scientific Officer of the AHAW unit
 Ticks and tick-borne pathogens Jordi Tarrés-Call, Scientific Officer of the AHAW unit Antwerp, June 2 nd 2010 1 The role of EFSA! To assess and communicate all risks associated with the food chain! We
Ticks and tick-borne pathogens Jordi Tarrés-Call, Scientific Officer of the AHAW unit Antwerp, June 2 nd 2010 1 The role of EFSA! To assess and communicate all risks associated with the food chain! We
Description of Culicoides (Culicoides) bysta n. sp., a new member of the Pulicaris group (Diptera: Ceratopogonidae) from Slovakia
 Sarvašová et al. Parasites & Vectors (2017) 10:279 DOI 10.1186/s13071-017-2195-4 RESEARCH Open Access Description of Culicoides (Culicoides) bysta n. sp., a new member of the Pulicaris group (Diptera:
Sarvašová et al. Parasites & Vectors (2017) 10:279 DOI 10.1186/s13071-017-2195-4 RESEARCH Open Access Description of Culicoides (Culicoides) bysta n. sp., a new member of the Pulicaris group (Diptera:
Summary of the latest data on antibiotic consumption in the European Union
 Summary of the latest data on antibiotic consumption in the European Union ESAC-Net surveillance data November 2016 Provision of reliable and comparable national antimicrobial consumption data is a prerequisite
Summary of the latest data on antibiotic consumption in the European Union ESAC-Net surveillance data November 2016 Provision of reliable and comparable national antimicrobial consumption data is a prerequisite
The OIE Manual of Diagnostic Tests and Vaccines for Terrestrial & Aquatic Animals
 The OIE Manual of Diagnostic Tests and Vaccines for Terrestrial & Aquatic Animals Regional seminar for OIE National Focal Points for Veterinary Products, Tokyo, Japan, 3-5 December 2014 Barbara Freischem,
The OIE Manual of Diagnostic Tests and Vaccines for Terrestrial & Aquatic Animals Regional seminar for OIE National Focal Points for Veterinary Products, Tokyo, Japan, 3-5 December 2014 Barbara Freischem,
Introduction to phylogenetic trees and tree-thinking Copyright 2005, D. A. Baum (Free use for non-commercial educational pruposes)
 Introduction to phylogenetic trees and tree-thinking Copyright 2005, D. A. Baum (Free use for non-commercial educational pruposes) Phylogenetics is the study of the relationships of organisms to each other.
Introduction to phylogenetic trees and tree-thinking Copyright 2005, D. A. Baum (Free use for non-commercial educational pruposes) Phylogenetics is the study of the relationships of organisms to each other.
Supplemental Information. Discovery of Reactive Microbiota-Derived. Metabolites that Inhibit Host Proteases
 Cell, Volume 168 Supplemental Information Discovery of Reactive Microbiota-Derived Metabolites that Inhibit Host Proteases Chun-Jun Guo, Fang-Yuan Chang, Thomas P. Wyche, Keriann M. Backus, Timothy M.
Cell, Volume 168 Supplemental Information Discovery of Reactive Microbiota-Derived Metabolites that Inhibit Host Proteases Chun-Jun Guo, Fang-Yuan Chang, Thomas P. Wyche, Keriann M. Backus, Timothy M.
Antimicrobial resistance (EARS-Net)
 SURVEILLANCE REPORT Annual Epidemiological Report for 2014 Antimicrobial resistance (EARS-Net) Key facts Over the last four years (2011 to 2014), the percentages of Klebsiella pneumoniae resistant to fluoroquinolones,
SURVEILLANCE REPORT Annual Epidemiological Report for 2014 Antimicrobial resistance (EARS-Net) Key facts Over the last four years (2011 to 2014), the percentages of Klebsiella pneumoniae resistant to fluoroquinolones,
Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST
 Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST INVESTIGATION 3 BIG IDEA 1 Lab Investigation 3: BLAST Pre-Lab Essential Question: How can bioinformatics be used as a tool to
Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST INVESTIGATION 3 BIG IDEA 1 Lab Investigation 3: BLAST Pre-Lab Essential Question: How can bioinformatics be used as a tool to
UW College of Agriculture and Natural Resources Global Perspectives Grant Program Project Report
 UW College of Agriculture and Natural Resources Global Perspectives Grant Program Project Report COVER PAGE Award Period: Fall 2017 Fall 2018 Principle Investigator: Brant Schumaker Department: Veterinary
UW College of Agriculture and Natural Resources Global Perspectives Grant Program Project Report COVER PAGE Award Period: Fall 2017 Fall 2018 Principle Investigator: Brant Schumaker Department: Veterinary
OIE Reference Laboratory Reports Activities
 OIE Reference Laboratory Reports Activities Activities in 2013 This report has been submitted : 2014-01-31 10:09:49 Name of disease (or topic) for which you are a designated OIE Reference Laboratory: Rabies
OIE Reference Laboratory Reports Activities Activities in 2013 This report has been submitted : 2014-01-31 10:09:49 Name of disease (or topic) for which you are a designated OIE Reference Laboratory: Rabies
Comparing DNA Sequences Cladogram Practice
 Name Period Assignment # See lecture questions 75, 122-123, 127, 137 Comparing DNA Sequences Cladogram Practice BACKGROUND Between 1990 2003, scientists working on an international research project known
Name Period Assignment # See lecture questions 75, 122-123, 127, 137 Comparing DNA Sequences Cladogram Practice BACKGROUND Between 1990 2003, scientists working on an international research project known
EFSA Scientific Opinion on canine leishmaniosis
 EFSA Scientific Opinion on canine leishmaniosis Andrea Gervelmeyer Animal Health and Welfare Team Animal and Plant Health Unit AHAC meeting 19 June 2015 PRESENTATION OUTLINE Outline Background ToR Approach
EFSA Scientific Opinion on canine leishmaniosis Andrea Gervelmeyer Animal Health and Welfare Team Animal and Plant Health Unit AHAC meeting 19 June 2015 PRESENTATION OUTLINE Outline Background ToR Approach
Ch 1.2 Determining How Species Are Related.notebook February 06, 2018
 Name 3 "Big Ideas" from our last notebook lecture: * * * 1 WDYR? Of the following organisms, which is the closest relative of the "Snowy Owl" (Bubo scandiacus)? a) barn owl (Tyto alba) b) saw whet owl
Name 3 "Big Ideas" from our last notebook lecture: * * * 1 WDYR? Of the following organisms, which is the closest relative of the "Snowy Owl" (Bubo scandiacus)? a) barn owl (Tyto alba) b) saw whet owl
Structure of the OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals
 Structure of the OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals Regional Seminar for OIE National Focal Points for Veterinary Laboratories Jeju, Republic of Korea, 5-7 April 2016 Dr.
Structure of the OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals Regional Seminar for OIE National Focal Points for Veterinary Laboratories Jeju, Republic of Korea, 5-7 April 2016 Dr.
COMMISSION DELEGATED REGULATION (EU)
 L 296/6 Official Journal of the European Union 15.11.2011 COMMISSION DELEGATED REGULATION (EU) No 1152/2011 of 14 July 2011 supplementing Regulation (EC) No 998/2003 of the European Parliament and of the
L 296/6 Official Journal of the European Union 15.11.2011 COMMISSION DELEGATED REGULATION (EU) No 1152/2011 of 14 July 2011 supplementing Regulation (EC) No 998/2003 of the European Parliament and of the
Finnzymes Oy. PathoProof Mastitis PCR Assay. Real time PCR based mastitis testing in milk monitoring programs
 PathoProof TM Mastitis PCR Assay Mikko Koskinen, Ph.D. Director, Diagnostics, Finnzymes Oy Real time PCR based mastitis testing in milk monitoring programs PathoProof Mastitis PCR Assay Comparison of the
PathoProof TM Mastitis PCR Assay Mikko Koskinen, Ph.D. Director, Diagnostics, Finnzymes Oy Real time PCR based mastitis testing in milk monitoring programs PathoProof Mastitis PCR Assay Comparison of the
MRSA surveillance 2014: Poultry
 Vicky Jasson MRSA surveillance 2014: Poultry 1. Introduction In the framework of the FASFC surveillance, a surveillance of MRSA in poultry has been executed in order to determine the prevalence and diversity
Vicky Jasson MRSA surveillance 2014: Poultry 1. Introduction In the framework of the FASFC surveillance, a surveillance of MRSA in poultry has been executed in order to determine the prevalence and diversity
Development and improvement of diagnostics to improve use of antibiotics and alternatives to antibiotics
 Priority Topic B Diagnostics Development and improvement of diagnostics to improve use of antibiotics and alternatives to antibiotics The overarching goal of this priority topic is to stimulate the design,
Priority Topic B Diagnostics Development and improvement of diagnostics to improve use of antibiotics and alternatives to antibiotics The overarching goal of this priority topic is to stimulate the design,
OIE Collaborating Centres Reports Activities
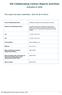 OIE Collaborating Centres Reports Activities Activities in 2016 This report has been submitted : 2017-01-20 17:44:12 Title of collaborating centre: Maladies infectieuses de la reproduction en Europe Address
OIE Collaborating Centres Reports Activities Activities in 2016 This report has been submitted : 2017-01-20 17:44:12 Title of collaborating centre: Maladies infectieuses de la reproduction en Europe Address
Animal reservoirs for Nipah virus
 Animal reservoirs for Nipah virus Dr. D. T. Mourya ICMR-National Institute of Virology Pune 411021, INDIA Tracing the source of Infection ICMR-NIV, Pune has team of scientific experts and trained field
Animal reservoirs for Nipah virus Dr. D. T. Mourya ICMR-National Institute of Virology Pune 411021, INDIA Tracing the source of Infection ICMR-NIV, Pune has team of scientific experts and trained field
Required and Recommended Supporting Information for IUCN Red List Assessments
 Required and Recommended Supporting Information for IUCN Red List Assessments This is Annex 1 of the Rules of Procedure for IUCN Red List Assessments 2017 2020 as approved by the IUCN SSC Steering Committee
Required and Recommended Supporting Information for IUCN Red List Assessments This is Annex 1 of the Rules of Procedure for IUCN Red List Assessments 2017 2020 as approved by the IUCN SSC Steering Committee
Can insecticide-treated netting provide protection for Equids from Culicoides biting midges in the United Kingdom?
 Baker et al. Parasites & Vectors (2015) 8:604 DOI 10.1186/s13071-015-1182-x RESEARCH Can insecticide-treated netting provide protection for Equids from Culicoides biting midges in the United Kingdom? Open
Baker et al. Parasites & Vectors (2015) 8:604 DOI 10.1186/s13071-015-1182-x RESEARCH Can insecticide-treated netting provide protection for Equids from Culicoides biting midges in the United Kingdom? Open
Bioinformatics: Investigating Molecular/Biochemical Evidence for Evolution
 Bioinformatics: Investigating Molecular/Biochemical Evidence for Evolution Background How does an evolutionary biologist decide how closely related two different species are? The simplest way is to compare
Bioinformatics: Investigating Molecular/Biochemical Evidence for Evolution Background How does an evolutionary biologist decide how closely related two different species are? The simplest way is to compare
RICKETTSIA SPECIES AMONG TICKS IN AN AREA OF JAPAN ENDEMIC FOR JAPANESE SPOTTED FEVER
 RICKETTSIA SPECIES AMONG TICKS IN AN AREA OF JAPAN ENDEMIC FOR JAPANESE SPOTTED FEVER Makoto Kondo 1, Katsuhiko Ando 2, Keiichi Yamanaka 1 and Hitoshi Mizutani 1 1 Department of Dermatology, 2 Department
RICKETTSIA SPECIES AMONG TICKS IN AN AREA OF JAPAN ENDEMIC FOR JAPANESE SPOTTED FEVER Makoto Kondo 1, Katsuhiko Ando 2, Keiichi Yamanaka 1 and Hitoshi Mizutani 1 1 Department of Dermatology, 2 Department
OIE Reference Laboratory Reports Activities
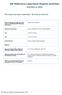 OIE Reference Laboratory Reports Activities Activities in 2016 This report has been submitted : 2017-01-13 10:41:13 Name of disease (or topic) for which you are a designated OIE Reference Laboratory: Enzootic
OIE Reference Laboratory Reports Activities Activities in 2016 This report has been submitted : 2017-01-13 10:41:13 Name of disease (or topic) for which you are a designated OIE Reference Laboratory: Enzootic
International Training Programme on Bluetongue Vector Identification (Meeting Book)
 International Training Programme on Bluetongue Vector Identification (Meeting Book) 1 st to 5 th July, 2013 Organized by The Pirbright Institute, UK India Bluetongue Vector Network && Tamil Nadu Veterinary
International Training Programme on Bluetongue Vector Identification (Meeting Book) 1 st to 5 th July, 2013 Organized by The Pirbright Institute, UK India Bluetongue Vector Network && Tamil Nadu Veterinary
PARTIAL REPORT. Juvenile hybrid turtles along the Brazilian coast RIO GRANDE FEDERAL UNIVERSITY
 RIO GRANDE FEDERAL UNIVERSITY OCEANOGRAPHY INSTITUTE MARINE MOLECULAR ECOLOGY LABORATORY PARTIAL REPORT Juvenile hybrid turtles along the Brazilian coast PROJECT LEADER: MAIRA PROIETTI PROFESSOR, OCEANOGRAPHY
RIO GRANDE FEDERAL UNIVERSITY OCEANOGRAPHY INSTITUTE MARINE MOLECULAR ECOLOGY LABORATORY PARTIAL REPORT Juvenile hybrid turtles along the Brazilian coast PROJECT LEADER: MAIRA PROIETTI PROFESSOR, OCEANOGRAPHY
Mosquitoes in a changing environment
 Mosquitoes in a changing environment Anders Lindström National Veterinary Institute Sweden Tree hole mosquito, Aedes geniculatus The One health concept is the realization that we are connected to our environment
Mosquitoes in a changing environment Anders Lindström National Veterinary Institute Sweden Tree hole mosquito, Aedes geniculatus The One health concept is the realization that we are connected to our environment
OIE Reference Laboratory Reports Activities
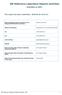 OIE Reference Laboratory Reports Activities Activities in 2017 This report has been submitted : 2018-01-24 10:31:11 Name of disease (or topic) for which you are a designated OIE Reference Laboratory: Classical
OIE Reference Laboratory Reports Activities Activities in 2017 This report has been submitted : 2018-01-24 10:31:11 Name of disease (or topic) for which you are a designated OIE Reference Laboratory: Classical
COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST
 COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST In this laboratory investigation, you will use BLAST to compare several genes, and then use the information to construct a cladogram.
COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST In this laboratory investigation, you will use BLAST to compare several genes, and then use the information to construct a cladogram.
Relative effectiveness of Irish factories in the surveillance of slaughtered cattle for visible lesions of tuberculosis,
 Iris Tréidliachta Éireann SHORT REPORT Open Access Relative effectiveness of Irish factories in the surveillance of slaughtered cattle for visible lesions of tuberculosis, 2005-2007 Francisco Olea-Popelka
Iris Tréidliachta Éireann SHORT REPORT Open Access Relative effectiveness of Irish factories in the surveillance of slaughtered cattle for visible lesions of tuberculosis, 2005-2007 Francisco Olea-Popelka
A Unique Approach to Managing the Problem of Antibiotic Resistance
 A Unique Approach to Managing the Problem of Antibiotic Resistance By: Heather Storteboom and Sung-Chul Kim Department of Civil and Environmental Engineering Colorado State University A Quick Review The
A Unique Approach to Managing the Problem of Antibiotic Resistance By: Heather Storteboom and Sung-Chul Kim Department of Civil and Environmental Engineering Colorado State University A Quick Review The
INQUIRY & INVESTIGATION
 INQUIRY & INVESTIGTION Phylogenies & Tree-Thinking D VID. UM SUSN OFFNER character a trait or feature that varies among a set of taxa (e.g., hair color) character-state a variant of a character that occurs
INQUIRY & INVESTIGTION Phylogenies & Tree-Thinking D VID. UM SUSN OFFNER character a trait or feature that varies among a set of taxa (e.g., hair color) character-state a variant of a character that occurs
The phenology and population dynamics of Culicoides spp. in different ecosystems in The Netherlands
 Available online at www.sciencedirect.com Preventive Veterinary Medicine 87 (2008) 41 54 www.elsevier.com/locate/prevetmed The phenology and population dynamics of Culicoides spp. in different ecosystems
Available online at www.sciencedirect.com Preventive Veterinary Medicine 87 (2008) 41 54 www.elsevier.com/locate/prevetmed The phenology and population dynamics of Culicoides spp. in different ecosystems
White Rose Research Online URL for this paper:
 This is an author produced version of Non-cultured faecal and gastrointestinal seed samples fail to detect Trichomonad infection in clinically and sub-clinically infected columbid birds. White Rose Research
This is an author produced version of Non-cultured faecal and gastrointestinal seed samples fail to detect Trichomonad infection in clinically and sub-clinically infected columbid birds. White Rose Research
NEOH Workshop on Evaluation of Data & Information Sharing in One Health Initiatives Copenhagen, 20 th & 21 st April 2016
 NEOH Workshop on Evaluation of Data & Information Sharing in One Health Initiatives Copenhagen, 20 th & 21 st April 2016 Prepare, Predict, Prevent: Creating Objectivity in Infectious Disease Risk Assessment
NEOH Workshop on Evaluation of Data & Information Sharing in One Health Initiatives Copenhagen, 20 th & 21 st April 2016 Prepare, Predict, Prevent: Creating Objectivity in Infectious Disease Risk Assessment
CERTIFIED REFERENCE MATERIAL IRMM 313
 EUROPEAN COMMISSION JOINT RESEARCH CENTRE Institute for Reference Materials and Measurements (Geel) CERTIFIED REFERENCE MATERIAL IRMM 313 CERTIFICATE OF ANALYSIS PFGE AGAROSE PLUGS Certified value 2) SmaI
EUROPEAN COMMISSION JOINT RESEARCH CENTRE Institute for Reference Materials and Measurements (Geel) CERTIFIED REFERENCE MATERIAL IRMM 313 CERTIFICATE OF ANALYSIS PFGE AGAROSE PLUGS Certified value 2) SmaI
The effects of diet upon pupal development and cocoon formation by the cat flea (Siphonaptera: Pulicidae)
 June, 2002 Journal of Vector Ecology 39 The effects of diet upon pupal development and cocoon formation by the cat flea (Siphonaptera: Pulicidae) W. Lawrence and L. D. Foil Department of Entomology, Louisiana
June, 2002 Journal of Vector Ecology 39 The effects of diet upon pupal development and cocoon formation by the cat flea (Siphonaptera: Pulicidae) W. Lawrence and L. D. Foil Department of Entomology, Louisiana
SCIENTIFIC REPORT. Analysis of the baseline survey on the prevalence of Salmonella in turkey flocks, in the EU,
 The EFSA Journal / EFSA Scientific Report (28) 198, 1-224 SCIENTIFIC REPORT Analysis of the baseline survey on the prevalence of Salmonella in turkey flocks, in the EU, 26-27 Part B: factors related to
The EFSA Journal / EFSA Scientific Report (28) 198, 1-224 SCIENTIFIC REPORT Analysis of the baseline survey on the prevalence of Salmonella in turkey flocks, in the EU, 26-27 Part B: factors related to
Drive More Efficient Clinical Action by Streamlining the Interpretation of Test Results
 White Paper: Templated Report Comments Drive More Efficient Clinical Action by Streamlining the Interpretation of Test Results Background The availability of rapid, multiplexed technologies for the comprehensive
White Paper: Templated Report Comments Drive More Efficient Clinical Action by Streamlining the Interpretation of Test Results Background The availability of rapid, multiplexed technologies for the comprehensive
