MORPHOMETRICS AND CYTOGENETICS OF Gracilinanus agilis AND Cryptonanus spp. (DIDELPHIMORPHIA: DIDELPHIDAE) FROM CENTRAL AND NORTHEASTERN BRAZIL
|
|
|
- Julian Gallagher
- 6 years ago
- Views:
Transcription
1 Mastozoología Neotropical, 17(1):53-60, Mendoza, 2010 SAREM, 2010 ISSN Versión on-line ISSN MORPHOMETRICS AND CYTOGENETICS OF Gracilinanus agilis AND Cryptonanus spp. (DIDELPHIMORPHIA: DIDELPHIDAE) FROM CENTRAL AND NORTHEASTERN BRAZIL João P. Garcia 1,3, João A. Oliveira 2, Margaret M. O. Corrêa 1, and Leila M. Pessôa 1 1 Laboratório de Mastozoologia, Departamento de Zoologia, Instituto de Biologia, Universidade Federal do Rio de Janeiro, Av. Brigadeiro Trompowski s/n, CCS, bl. A, sala A1-121, Ilha do Fundão, CEP: Rio de Janeiro, RJ Brasil [Corresponding author: João Pedro Garcia <araujo_jpg@yahoo.com.br>]. 2 Setor de Mamíferos, Departamento de Vertebrados, Museu Nacional/UFRJ, Quinta da Boa Vista, São Cristóvão, CEP: Rio de Janeiro, RJ Brasil. 3 Programa de Pós-Graduação em Ciências Biológicas (Zoologia), Museu Nacional/UFRJ, Brasil. ABSTRACT: Species of the didelphid genera Gracilinanus and Cryptonanus are morphologically and cytogenetically very similar. Several qualitative characters, some of which exhibit intraspecific polymorphisms, have been used to distinguish these genera, but more data are needed to characterize them better. Samples of G. agilis and Cryptonanus spp. from nine localities in central and northeastern Brazil were analyzed. Multivariate analyses of craniodental measurements and descriptive statistics of external body measurements indicate that G. agilis is conspicuously larger than Cryptonanus spp., and that general size is the main factor distinguishing these forms. Size differences can be combined with qualitative characters making the differentiation between G. agilis and Cryptonanus spp. easier. Cytogenetic analyses, including the first description of C-bands and Ag-NORs of G. agilis, revealed that the karyotypes of G. agilis and Cryptonanus sp. from Barão de Melgaço, Mato Grosso, are very similar, except for the fourth autosomal pair and the X chromosome. RESUMEN: Morfometría y citogenética de Gracilinanus agilis y Cryptonanus spp. (Didelphimorphia: Didelphidae) del centro y nordeste del Brasil. Los didélfidos Gracilinanus y Cryptonanus poseen una morfología y citogenética muy semejantes. Estos géneros han sido diferenciados por caracteres polimórficos cualitativos, pero más datos son necesarios para caracterizarlos mejor. Fueron analizadas muestras de G. agilis y Cryptonanus spp. de nueve localidades de centro y nordeste de Brasil. Los análisis multivariados de las medidas craneodentarias y las estadísticas descriptivas de las medidas corporales externas indican que G. agilis es claramente mayor que Cryptonanus spp. y que el tamaño general es el principal factor para distinguir esas formas. La variación en tamaño puede ser asociada a los caracteres cualitativos para facilitar la diferenciación entre G. agilis y Cryptonanus spp. Los análisis citogenéticos, incluyendo la primera descripción del bandeo C y las Ag-NORs de G. agilis, revelaron que los cariotipos de G. agilis y Cryptonanus sp. de Barão de Melgaço, Mato Grosso, son muy semejantes, excepto por el cuarto par de autosomas y el cromosoma X. Key words. Differentiation. General size. Karyotypes. Mouse opossums. Palabras clave. Cariotipos. Diferenciación. Marmosinos. Tamaño general. Recibido 11 marzo Aceptado 18 julio Editor asociado: G D Elía
2 54 Mastozoología Neotropical, 17(1):53-60, Mendoza, 2010 JP Garcia et al. INTRODUCTION The genus Gracilinanus Gardner and Creighton, 1989, as recently restricted by Voss et al. (2005) comprises six species: G. aceramarcae (Tate, 1931), G. agilis (Burmeister, 1854), G. dryas (Thomas, 1898), G. emiliae (Thomas, 1909), G. marica (Thomas, 1898) and G. microtarsus (Wagner, 1842). These are long-tailed, small-sized (head and body, mm; tail, mm; weight < 50g), pouchless opossums, with a dark circumocular mask (Gardner and Creighton, 1989; Voss et al., 2005). The dorsal pelage ranges from bright reddish brown to dull brownish gray and the ventral pelage ranges from white to pale orange with gray-based hairs present to a greater or lesser extent. The tail ranges from moderately long to very long and can be unicolored fuscous or weakly bicolored through its length. The presence of maxillary fenestrae and of a secondary foramen ovale formed by the anteromedial bullar process are considered two of the most important diagnostic cranial characters for the genus (Costa et al., 2003). Before the genus Cryptonanus was described by Voss et al. (2005), the forms now recognized as C. agricolai (Moojen, 1943), C. chacoensis (Tate, 1931), C. guahybae (Tate, 1931), C. ignitus (Díaz, Flores and Barquez, 2002), and C. unduaviensis (Tate, 1931) were included within Gracilinanus. Although species of Gracilinanus and Cryptonanus are considered very similar, Voss et al. (2005) described some discrete morphological characters that could be used to distinguish these genera. In spite of some polymorphisms, most species of Cryptonanus lack a rostral process in the premaxillae, a secondary foramen ovale, and maxillary fenestrae; the second upper premolar is shorter than the third one; and the upper canine has accessory cusps. Besides these characters, these authors also suggested that Cryptonanus specimens have shorter rostrums and smaller orbits, but they remarked that these proportions are ontogenetically variable and that available samples were too small for statistically compelling morphometric analyses. Cytogenetic data on Gracilinanus and Cryptonanus are scarce. The karyotypes of G. microtarsus and C. chacoensis from Argentina (the latter identified as Marmosa agilis chacoensis) were described by Wainberg et al. (1979). These authors recorded the diploid number (2n) and described the chromosomal morphology for both species. Carvalho et al. (2002) described the 2n and the number of autosomal arms (FN) of G. agilis and C. agricolai (the latter identified as G. emiliae), and the distribution of both the constitutive heterochromatin (C-bands) and the silver stained nucleolar organizer regions (Ag-NORs) in G. microtarsus and C. agricolai. All of these species showed 2n = 14 and FN = 24. The karyotypes of the species listed above, except C. chacoensis, were described on the basis of Brazilian specimens. Three species of Gracilinanus are known to occur in Brazil: G. emiliae, known from two localities in Pará state (Voss et al., 2001); G. microtarsus, known from mesic habitats of the Atlantic Forest; and G. agilis, known from dry forests and gallery forests of central Brazil (Costa et al., 2003). Three species of Cryptonanus have also been recorded for Brazil: C. chacoensis, known from northern Pantanal (Rossi et al., 2006); C. agricolai, known from the Caatinga and Cerrado of eastcentral Brazil; and C. guahybae, known from a few localities in the state of Rio Grande do Sul (Voss et al., 2005). Two areas of sympatry are known between species of Gracilinanus and Cryptonanus in Brazil. Gracilinanus microtarsus is sympatric with C. guahybae in Rio Grande do Sul and G. agilis is sympatric with C. agricolai and C. chacoensis in central and northeastern Brazil. Herein we evaluate the morphometric and cytogenetic variation in Gracilinanus agilis, the smallest species of this genus (Costa et al., 2003), which is known to occur sympatrically with species of Cryptonanus. Considering that most of the qualitative morphological characters previously used to differentiate these gen-
3 Gracilinanus agilis AND Cryptonanus spp. FROM BRAZIL 55 era are polymorphic, we intend to evaluate if morphometric data can be used to distinguish G. agilis from Cryptonanus spp. We complement the characterization of these taxa with cytogenetic data, including the first description of C-bands and Ag-NORs of G. agilis and the karyotype with Ag-NORs of Cryptonanus sp. from Barão de Melgaço, Mato Grosso. MATERIAL AND METHODS We analyzed 59 specimens of G. agilis and 8 specimens of Cryptonanus spp. from 9 localities in central and northeastern Brazil (Fig. 1). Based on provisional diagnoses given in Voss et al. (2005), specimens of Cryptonanus present in our restricted samples could be assigned to the nominal forms agricolai (specimens from Goiás, Bahia and Ceará) and chacoensis (specimens from Mato Grosso). However, due to the generic level of the comparisons made here, we treated these forms as Cryptonanus spp., to avoid misidentifications caused by the relatively unclear status of these forms diagnoses (Voss et al., 2005). Thirteen specimens of G. agilis and two of Cryptonanus sp. from Barão de Melgaço, Mato Grosso, were karyotyped. More data on localities and specimens are presented in Appendix. Fig. 1. Collection sites of specimens of Gracilinanus agilis and Cryptonanus spp. in central and northeastern Brazil. State of Mato Grosso do Sul: Corumbá (1) and Aquidauana (2); State of Mato Grosso: Barão de Melgaço (3); State of Goiás: Anápolis (4), Serra da Mesa (5), Teresina de Goiás (6), and Cavalcante (7); State of Bahia: Chapada Diamantina (8); State of Ceará: Crato (9). We recorded 20 cranial and dental measurements with digital calipers accurate to 0.01 mm. Definitions of the following characters can be found in Hershkovitz (1992), Costa et al. (2003), and Voss et al. (2005): greatest skull length (GSL), condylobasal length (CBL), basal skull length (BSL), rostral length (ROL), nasal length (NAL), braincase breadth (BCB), zygomatic breadth (ZYB), postorbital breadth (POB), interorbital breadth (IOB), rostral breadth (ROB), cranial depth (CRD), length between first incisor and last molar (IM4), length from canine to last molar (CM4), length from first to last molar (MM4), length from first to third molar (MM3), palatal length (PAL), palatal breadth (PAB), least pterygoid breadth (LPB), petrosal bulla breadth (PBB), and alisphenoid bulla breadth (ABB). In addition to these measurements, we also recorded mandibular length (MDL, measured from the anterior extremity of the dentarium to the condyloid process). We recorded the following external measurements from museum skin tags: head and body length (HB), length of tail (LT), length of hind foot (including the claw, HF), and length of ear (Ear). All measurements, craniodental and external, were taken from adult specimens (age classes 6 and 7 of Tribe, 1990). Significant sexual dimorphism was detected within samples of G. agilis, but unfortunately we could not test this dimorphism in Cryptonanus spp. because our restricted sample contains only one female. Males and females of both taxa were grouped in all analyses because we presumed that intergeneric differences should be greater than intraspecific differences. Descriptive statistics for external measurements were calculated. All craniodental measurements were log-transformed for multivariate statistical analyses. Missing values (2.4%) were estimated by the expectation-maximization method (Strauss et al., 2003). A principal component analysis was employed in the variance-covariance matrix to search of the main patterns of craniometric variation in the total dataset including available samples of adult specimens of G. agilis and Cryptonanus spp. Character correlations with principal components were portrayed as vector plots. Multivariate statistical analyses were performed on MatLab 4.3 (The MathWorks) using routines written by R. Strauss (available at Strauss/Matlab/matlab.htm). We employed the technique of Ford and Hamerton (1956) for mitotic preparations. C-banding and Ag-NOR sites were detected employing the techniques of Sumner (1972) and Howell and
4 56 Mastozoología Neotropical, 17(1):53-60, Mendoza, 2010 JP Garcia et al. Black (1980), respectively. The chromosomal nomenclature follows Patton (1967). RESULTS Principal Component Analysis revealed that a single axis (PC1) accounts for most of the variance (approximately 82%) of craniodental measurement data. Two major clusters were detected along the first axis: one cluster comprises G. agilis samples and the other comprises samples referred to Cryptonanus spp. (Fig. 2). Most characters are positively associated with the PC1, indicating that it can be interpreted as a general size vector (Fig. 3). The second axis (PC2) accounts for approximately 6% of the variance, while the other 12% are distributed over the third to the fifth axes. Along the second axis, the local samples that compose the cluster of G. agilis are notably overlapped, whereas the local samples of Cryptonanus cluster show no superposition. The sample from Crato, composed by the type and paratype of C. agricolai, does not overlap with the remaining samples referred to Cryptonanus, which are distributed along PC2 (Fig. 2). Least pterygoid breadth and postorbital breadth are the characters that best separate the subgroups of Cryptonanus (Fig. 3). Analysis of external measurements reveals overlap between the observed range of variation in G. agilis and Cryptonanus spp. for HB, LT, Ear, and HF. Nevertheless, for all external measurements mean values of G. agilis are higher than those of Cryptonanus spp. (Table 1). Cytogenetic analyses of specimens of G. agilis from Barão de Melgaço, Mato Grosso, showed 2n = 14 and FN = 24, with six pairs of biarmed autosomes. Pairs 1, 2 and 3 are submetacentric; pair 4 is metacentric; and pairs 5 and 6 are subtelocentric. Sex chromosomes are the smallest pair in the karyotype. The X chromosome is submetacentric and the Y is acrocentric (Fig. 4A). Analyses of C-banding patterns showed small blocks of constitutive heterochromatin located at the pericentromeric regions of all autosomes and the X chromosome. The Y chromosome is entirely hetero- Fig. 2. First and second components of principal component analysis of 21 craniodental measurements of Gracilinanus agilis (1-6) and Cryptonanus spp. (7-10) from central and northeastern Brazil. Localities are Barão de Melgaço (1,7), Cavalcante (2), Teresina de Goiás (3), Aquidauana + Corumbá (4), Chapada Diamantina (5, 8), Serra da Mesa (6), Serra da Mesa + Anápolis (9), and Crato (10, type series of C. agricolai). Fig. 3. Vector plot of loadings of first and second components of the principal component analysis of 21 craniodental measurements of Gracilinanus agilis and Cryptonanus spp. from several localities from central and northeastern Brazil. POB postorbital breadth; LPB least pterygoid breadth; MM3 length from first to third molar; MM4 length from first to fourth molar; ROB rostral breadth. Some measurement acronyms are omitted.
5 Gracilinanus agilis AND Cryptonanus spp. FROM BRAZIL 57 Table 1 Descriptive statistics of external measurements of Gracilinanus agilis and Cryptonanus spp. from central and northeastern Brazil. G. agilis (n=38) Cryptonanus spp. (n=6) X Min Max X Min Max HB LT HF 18 a Ear 21 b c a n = 37; b n = 35; c n = 3. chromatic (Fig. 4B). The analysis of Ag-NORs revealed that these regions are only present on the short arm of autosome pair 6 (Fig. 4C). Cryptonanus sp. from Barão de Melgaço, Mato Grosso, showed 2n = 14 and NF = 24, with six pairs of biarmed autosomes. Pairs 1, 2, 3, and 4 are submetacentric; pairs 5 and 6 are subtelocentric. Sex chromosomes are the smallest pair in the karyotype. The X chromosome is subtelocentric (Fig. 5A). The analysis of Ag-NORs revealed that these regions are only present on the short arm of autosome pair 6 (Fig. 5B). DISCUSSION Multivariate analyses of craniodental traits revealed that adult samples of G. agilis and Cryptonanus spp. are clearly distinguishable based on general size. The interspecific differences are so conspicuous that intraspecific differences such as sexual dimorphism, which is known for G. agilis (see Costa et al., 2003), do not prevent species cluster formation by the analyses. Our results indicate that the differences between Cryptonanus spp. and G. agilis can be attributed to general cranial size and size-correlated (allometric) shape variation. Although there are conspicuous size differences between G. agilis and Cryptonanus spp., the identification of the latter becomes easier if qualitative characters pointed out by Voss et al. (2005) are combined with craniodental and body measurements. In spite of our restricted sample, it is possible to observe a difference between external measurement means of adults of G.agilis and Cryptonanus spp. However, juvenile, subadults, and even unusual small adults of G. agilis can be hard to distinguish from specimens of Cryptonanus spp. if only external measurements are considered. External quali- Fig. 4. Karyotype of Gracilinanus agilis from Barão de Melgaço, Mato Grosso, Brazil. A) Giemsa staining (MN64376); B) C-banding (MN64403); C) Ag-NORs (MN64403).
6 58 Mastozoología Neotropical, 17(1):53-60, Mendoza, 2010 JP Garcia et al. Fig. 5. Karyotype of Cryptonanus sp. from Barão de Melgaço, Mato Grosso, Brazil. A) Giemsa staining (MN64347); B) Ag-NORs (MN64260). tative traits as well as craniodental quantitative and qualitative traits must be used to guarantee the correct diagnoses of these taxa. Therefore, field identifications may not be attempted on the exclusive basis of external measurements. Although the aim of this paper was to evaluate the distinctiveness of G. agilis and Cryptonanus spp. based on morphometrics and cytogenetics, we believe that one qualitative character must be emphasized: one specimen of Cryptonanus from Anápolis, State of Goiás, presented gray-based ventral fur. According to distributional patterns presented by Voss et al. (2005) this specimen should be assigned to C. agricolai. In the provisional diagnosis of this species, all but one specimens presented self-whitish ventral fur (Voss et al., 2005); the specimen from Anápolis constitutes the second exception to this pattern. This corroborates the lack of knowledge about variation in Cryptonanus (see Voss et al., 2005), which is a consequence of the reduced number of specimens in museum collections. Regarding cytogenetic traits, G. agilis and Cryptonanus sp. from Barão de Melgaço, Mato Grosso, have very similar karyotypes, which is not unexpected because other didelphid species that have the same diploid number usually also have the same fundamental number and distribution pattern of Ag-NORs and C bands (Souza et al., 1990; Carvalho et al., 2002; Svartman and Vianna-Morgante, 2003). The two karyotypic differences between G. agilis and Cryptonanus sp. from Barão de Melgaço are in the fourth autosome pair, which is metacentric in the former and submetacentric in the latter, and in the X chromosome, which is submetacentric in the former and subtelocentric in the latter. The other autosomes and the position of Ag-NORs are identical in both species. We were not able to compare the Y chromosomes of both species. As this is the first description of the Ag- NORs and C-bands in G. agilis and of the karyotype of Cryptonanus sp. from Mato Grosso, we present a brief comparison of our data with literature records for other congeneric species. The pattern of Ag-NORs and C- bands distribution reported by us for G. agilis is the same as that described for G. microtarsus from the states of Santa Catarina and Rio Grande do Sul, Brazil (Carvalho et al., 2002). The pattern of Ag-NORs distribution we reported for Cryptonanus sp. from Barão de Melgaço is also identical to that described for specimens attributed by Voss et al. (2005) to C. agricolai from the State of Goiás, Brazil (Carvalho et al., 2002). These findings are in accord with the high degree of karyotypical conservatism previously attributed to didelphids (Reig et al., 1977; Souza et al., 1990; Carvalho et al., 2002; Svartman and Vianna-Morgante, 2003). A major concern in taxonomy of mouse opossums is the lack of knowledge on their
7 Gracilinanus agilis AND Cryptonanus spp. FROM BRAZIL 59 intraspecific variation, which is a consequence of the reduced number of specimens in museum collections. Increasing these samples is a key point to improve the understanding about the differences between G. agilis and Cryptonanus spp. Furthermore, this would enable us to shed more light on the provisional diagnoses of species of Cryptonanus given by Voss et al. (2005). ACKNWOLEDGEMENTS To Stella Maris Franco for technical support in Museu Nacional mammal collection and Paulo Sérgio D Andrea for the permission to study the specimens under his care. To A.L. Gomes for helping with the map. To Guillermo D Elía and an anonymous referee for the valuable suggestions to improve this manuscript. J.P. Garcia receives financial support from FAPERJ scholarship and L.M. Pessôa and J.A. Oliveira receive research grants from CNPq. LITERATURE CITED CARVALHO BA, LFB OLIVEIRA, AP NUNES, and MS MATTEVI Karyotypes of nineteen marsupial species from Brazil. Journal of Mammalogy 83: COSTA LP, YL LEITE, and JL PATTON Phylogeography and systematic notes on two species of gracile mouse opossums, genus Gracilinanus (Marsupialia: Didelphidae) from Brazil. Proceedings of the Biological Society of Washington 116: FORD CE and JL HAMERTON A colchicine, hypotonic citrate, squash sequence for mammalian chromosomes. Stain Technology 31: GARDNER AL and GK CREIGHTON A new generic name for Tate s microtarsus group of South American mouse opossums (Marsupialia: Didelphidae). Proceedings of the Biological Society of Washington 102:3-7. HERSHKOVITZ P The South American gracile mouse opossums, genus Gracilinanus Gardner & Creighton, 1989 (Marmosidae: Marsupialia): a taxonomic review with notes on general morphology and relationships. Fieldiana Zoology 86: HOWELL WM and DA BLACK Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. Experientia 36: PATTON JL Chromosome studies of certain pocket mice genus Perognathus (Rodentia: Heteromyidae). Journal of Mammalogy 48: REIG OA, AL GARDNER, NO BIANCHI, and JL PATTON The chromosome of the Didelphidae (Marsupialia) and their evolutionary significance. Biological Journal of Linnean Society 9: ROSSI RV, GV BIANCONI, and WA PEDRO Ordem Didelphimorphia. Pp , in: Mamíferos do Brasil (N Reis, A Peracchi, WA Pedro, and IP Lima, eds.). Londrina. SOKAL RR and FJ ROHLF Biometry. 3 rd Edition. W. H. Freeman and Company, New York. SOUZA MJ, V MAIA, and JF SANTOS Nucleolar organizer regions, G- and C-bands in some Brazilian species of didelphids. Revista Brasileira de Genética 13: STRAUSS RE, MN ATANASSOV, and JA OLIVEIRA Evaluation of the principal-component and expectation-maximization methods for estimating missing data in morphometric studies. Journal of Vertebrate Paleontology 23: SUMNER AT A simple technique for demonstrating centromeric heterocromatin. Experimental Cell Research 75: SVARTMAN M and AM VIANNA-MORGANTE Conservation of chromosomal location of nucleolus organizer in American marsupials (Didelphidae). Genetica 118: TRIBE CJ Dental age classes in Marmosa incana and other didelphoids. Journal of Mammalogy 71: VOSS RS, DP LUNDE, and SA JANSA On the contents of Gracilinanus Gardner & Creighton, 1989, with the description of a previously unrecognized clade of small didelphids marsupials. American Museum Novitates 3482:1-34. VOSS RS, DP LUNDE, and NB SIMMONS The mammals of Paracou, French Guiana: a Neotropical lowland rainforest fauna, part 2. Nonvolant species. Bulletin of American Museum of Natural History 263: WAINBERG RL, TG DE FRONZA, and JG GARCIA Cromosomas de marsupiales del género Marmosa: M. pusilla bruchi, M. agilis chacoensis y M. microtarsus (Marsupialia: Didelphidae). Physis 38:33-38.
8 60 Mastozoología Neotropical, 17(1):53-60, Mendoza, 2010 JP Garcia et al. APPENDIX All voucher specimens analyzed in this study are deposited in the Museu Nacional (MN), Rio de Janeiro, Brazil. The cell suspension samples (*) used in the chromosomic analysis are deposited in the Laboratório de Mastozoologia, Departamento de Zoologia IB/UFRJ. Cryptonanus spp. BAHIA: Chapada Diamantina (13º32 S, 41º50 W) MN CEARÁ: Crato (7º14 S, 39º24 W) MN1494, MN1495 (paratype and holotype of C. agricolai, respectively). GOIÁS: Anápolis (16º19 S, 48º57 W) MN50165; Serra da Mesa (14º12 S, 48º03 W) MN36216, MN MATO GROSSO: Barão de Melgaço (16 11 S, W) MN64260*, MN64347*. Gracilinanus agilis: BAHIA: Chapada Diamantina MN67582, MN67599, MN67606, MN67607, MN67611, MN GOIÁS: Cavalcante (13º47 S, 47º27 W) MN46538, MN46539, MN46540, MN46541, MN46542, MN46543, MN46544, MN46545, MN46546, MN46547, MN46548, MN46549, MN46550, MN46551, MN46552, MN46567; Serra da Mesa MN36023, MN36047, MN36085, MN36201, MN36279, MN36292, MN36735; Teresina de Goiás (13º46 S, 47º15 W) MN42979, MN42981, MN42982, MN MATO GROSSO: Barão de Melgaço MN64286*, MN64376*, MN64403*, MN64412*, MN64488*, MN64497*, MN64498*, MN64499*, MN64539*, MN64628*, MN64663*, MN64695*, MN64696*, MN MATO GROSSO DO SUL: Aquidauana (20º28 S, 55º47 W) LBCE4823, LBCE4824, LBCE4835, LBCE4840, LBCE4846, LBCE4854, LBCE4876, LBCE4888, LBCE6145, LBCE6149; Corumbá (19º00 S, 57º39 W) LBCE5691, LBCE5700, LBCE6147.
CHACO MOUSE OPOSSUM Cryptonanus chacoensis (Tate, 1931)
 CHACO MOUSE OPOSSUM Cryptonanus chacoensis (Tate, 1931) FIGURE 1 - (FPMAM14PH) Adult, Estancia Nueva Gambach, PN San Rafael, Departamento Itapúa (Flavia Netto December 2008). TAXONOMY: Class Mammalia;
CHACO MOUSE OPOSSUM Cryptonanus chacoensis (Tate, 1931) FIGURE 1 - (FPMAM14PH) Adult, Estancia Nueva Gambach, PN San Rafael, Departamento Itapúa (Flavia Netto December 2008). TAXONOMY: Class Mammalia;
A new karyotypic formula for the genus Amphisbaena (Squamata: Amphisbaenidae)
 Phyllomedusa 9(1):75-80, 2010 2010 Departamento de Ciências Biológicas - ESALQ - USP ISSN 1519-1397 Short Communication A new karyotypic formula for the genus Amphisbaena (Squamata: Amphisbaenidae) Camila
Phyllomedusa 9(1):75-80, 2010 2010 Departamento de Ciências Biológicas - ESALQ - USP ISSN 1519-1397 Short Communication A new karyotypic formula for the genus Amphisbaena (Squamata: Amphisbaenidae) Camila
AGILE GRACILE OPOSSUM Gracilinanus agilis (Burmeister, 1854)
 AGILE GRACILE OPOSSUM Gracilinanus agilis (Burmeister, 1854) FIGURE 1 - Adult, Brazil (Nilton Caceres undated). TAXONOMY: Class Mammalia; Subclass Theria; Infraclass Metatheria; Magnorder Ameridelphia;
AGILE GRACILE OPOSSUM Gracilinanus agilis (Burmeister, 1854) FIGURE 1 - Adult, Brazil (Nilton Caceres undated). TAXONOMY: Class Mammalia; Subclass Theria; Infraclass Metatheria; Magnorder Ameridelphia;
Key words. Amazonia. Mammal. Marsupial. Neotropical. Rainforest. Palabras claves. Amazonía. Mamífero. Marsupial. Neotropical. Selva.
 Mastozoología Neotropical, 16(2):433-443, Mendoza, 2009 SAREM, 2009 ISSN 0327-9383 Versión on-line ISSN 1666-0536 ON THE DIAGNOSTIC CHARACTERS, ECOGEOGRAPHIC DISTRIBUTION, AND PHYLOGENETIC RELATIONSHIPS
Mastozoología Neotropical, 16(2):433-443, Mendoza, 2009 SAREM, 2009 ISSN 0327-9383 Versión on-line ISSN 1666-0536 ON THE DIAGNOSTIC CHARACTERS, ECOGEOGRAPHIC DISTRIBUTION, AND PHYLOGENETIC RELATIONSHIPS
Reptilia, Squamata, Amphisbaenidae, Anops bilabialatus : Distribution extension, meristic data, and conservation.
 Reptilia, Squamata, Amphisbaenidae, Anops bilabialatus : Distribution extension, meristic data, and conservation. Tamí Mott 1 Drausio Honorio Morais 2 Ricardo Alexandre Kawashita-Ribeiro 3 1 Departamento
Reptilia, Squamata, Amphisbaenidae, Anops bilabialatus : Distribution extension, meristic data, and conservation. Tamí Mott 1 Drausio Honorio Morais 2 Ricardo Alexandre Kawashita-Ribeiro 3 1 Departamento
A NEW SPECIES OF GRACILE MOUSE OPOSSUM, GENUS GRACILINANUS (DIDELPHIMORPHIA: DIDELPHIDAE), FROM ARGENTINA
 Journal of Mammalogy, 8():82 8, 2002 A NEW SPECIES OF GRACILE MOUSE OPOSSUM, GENUS GRACILINANUS (DIDELPHIMORPHIA: DIDELPHIDAE), FROM ARGENTINA M. MÓNICA DíAZ,* DAVID A. FLORES, AND RUBÉN M. BARQUEZ Sam
Journal of Mammalogy, 8():82 8, 2002 A NEW SPECIES OF GRACILE MOUSE OPOSSUM, GENUS GRACILINANUS (DIDELPHIMORPHIA: DIDELPHIDAE), FROM ARGENTINA M. MÓNICA DíAZ,* DAVID A. FLORES, AND RUBÉN M. BARQUEZ Sam
FIRST RECORD OF Platemys platycephala melanonota ERNST,
 FIRST RECORD OF Platemys platycephala melanonota ERNST, 1984 (REPTILIA, TESTUDINES, CHELIDAE) FOR THE BRAZILIAN AMAZON Telêmaco Jason Mendes-Pinto 1,2 Sergio Marques de Souza 2 Richard Carl Vogt 2 Rafael
FIRST RECORD OF Platemys platycephala melanonota ERNST, 1984 (REPTILIA, TESTUDINES, CHELIDAE) FOR THE BRAZILIAN AMAZON Telêmaco Jason Mendes-Pinto 1,2 Sergio Marques de Souza 2 Richard Carl Vogt 2 Rafael
Karyotype, constitutive heterochromatin and nucleolus organizer regions in two species of Liolaemus (Squamata, Tropiduridae)
 CARYOLOGIA Vol. 56, no. 3: 269-273, 2003 Karyotype, constitutive heterochromatin and nucleolus organizer regions in two species of Liolaemus (Squamata, Tropiduridae) ALEJANDRA HERNANDO Departamento de
CARYOLOGIA Vol. 56, no. 3: 269-273, 2003 Karyotype, constitutive heterochromatin and nucleolus organizer regions in two species of Liolaemus (Squamata, Tropiduridae) ALEJANDRA HERNANDO Departamento de
On the Relationships of Marmosa formosa Shamel, 1930 (Marsupialia: Didelphidae), a Phylogenetic Puzzle from the Chaco of Northern Argentina
 PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, NY 10024 Number 3442, 18 pp., 6 figures, 4 tables June 2, 2004 On the Relationships of Marmosa formosa Shamel,
PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, NY 10024 Number 3442, 18 pp., 6 figures, 4 tables June 2, 2004 On the Relationships of Marmosa formosa Shamel,
BRAZILIAN GRACILE OPOSSUM Gracilinanus microtarsus (JA Wagner, 1842)
 BRAZILIAN GRACILE OPOSSUM Gracilinanus microtarsus (JA Wagner, 1842) FIGURE 1 - Adult, São Paulo, Brazil ( Thomas Püttker February 2005). TAXONOMY: Class Mammalia; Subclass Theria; Infraclass Metatheria;
BRAZILIAN GRACILE OPOSSUM Gracilinanus microtarsus (JA Wagner, 1842) FIGURE 1 - Adult, São Paulo, Brazil ( Thomas Püttker February 2005). TAXONOMY: Class Mammalia; Subclass Theria; Infraclass Metatheria;
Title: Phylogenetic Methods and Vertebrate Phylogeny
 Title: Phylogenetic Methods and Vertebrate Phylogeny Central Question: How can evolutionary relationships be determined objectively? Sub-questions: 1. What affect does the selection of the outgroup have
Title: Phylogenetic Methods and Vertebrate Phylogeny Central Question: How can evolutionary relationships be determined objectively? Sub-questions: 1. What affect does the selection of the outgroup have
What we ve covered so far:
 What we ve covered so far: Didelphimorphia Didelphidae opossums (1 B.C. species) Soricomorpha Soricidae shrews (9 B.C. species) Talpidae moles (3 B.C. species) What s next: Rodentia Sciuridae squirrels
What we ve covered so far: Didelphimorphia Didelphidae opossums (1 B.C. species) Soricomorpha Soricidae shrews (9 B.C. species) Talpidae moles (3 B.C. species) What s next: Rodentia Sciuridae squirrels
Taxonomy and distribution of the Brazilian species of Thylamys (Didelphimorphia: Didelphidae)
 Mammalia (2006): 126 144 2006 by Walter de Gruyter Berlin New York. DOI 10.1515/MAMM.2006.013 Taxonomy and distribution of the Brazilian species of Thylamys (Didelphimorphia: Didelphidae) La taxonomie
Mammalia (2006): 126 144 2006 by Walter de Gruyter Berlin New York. DOI 10.1515/MAMM.2006.013 Taxonomy and distribution of the Brazilian species of Thylamys (Didelphimorphia: Didelphidae) La taxonomie
Fig. 5. (A) Scaling of brain vault size (width measured at the level of anterior squamosal/parietal suture) relative to skull size (measured at the
 Fig. 5. (A) Scaling of brain vault size (width measured at the level of anterior squamosal/parietal suture) relative to skull size (measured at the distance between the left versus right temporomandibular
Fig. 5. (A) Scaling of brain vault size (width measured at the level of anterior squamosal/parietal suture) relative to skull size (measured at the distance between the left versus right temporomandibular
Mammalogy Lab 1: Skull, Teeth, and Terms
 Mammalogy Lab 1: Skull, Teeth, and Terms Be able to: Goals of today s lab Locate all structures listed on handout Define all terms on handout what they are or what they look like Give examples of mammals
Mammalogy Lab 1: Skull, Teeth, and Terms Be able to: Goals of today s lab Locate all structures listed on handout Define all terms on handout what they are or what they look like Give examples of mammals
Description of Malacomys verschureni, a new Murid-species from Central Africa
 (Rev. ZooI. afr., 91, no 3) (A paru Ie 30 septembre 1977). Description of Malacomys verschureni, a new Murid-species from Central Africa (Mammalia - Muridae) By W.N. VERHEYEN ANDE. VAN DER STRAETEN * (Antwerpen)
(Rev. ZooI. afr., 91, no 3) (A paru Ie 30 septembre 1977). Description of Malacomys verschureni, a new Murid-species from Central Africa (Mammalia - Muridae) By W.N. VERHEYEN ANDE. VAN DER STRAETEN * (Antwerpen)
A New Species of Chiroderma from Guadeloupe, West Indies (Chiroptera: Phyllostomidae)
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Mammalogy Papers: University of Nebraska State Museum Museum, University of Nebraska State 4-16-1976 A New Species of Chiroderma
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Mammalogy Papers: University of Nebraska State Museum Museum, University of Nebraska State 4-16-1976 A New Species of Chiroderma
Revista de Biología Tropical ISSN: Universidad de Costa Rica Costa Rica
 Revista de Biología Tropical ISSN: 0034-7744 rbt@cariari.ucr.ac.cr Universidad de Costa Rica Costa Rica Gonçalves, Inês C.; Da-Silva, Elidiomar R.; Nessimian, Jorge L. Oligoneuria macabaiba sp. nov. (Insecta:
Revista de Biología Tropical ISSN: 0034-7744 rbt@cariari.ucr.ac.cr Universidad de Costa Rica Costa Rica Gonçalves, Inês C.; Da-Silva, Elidiomar R.; Nessimian, Jorge L. Oligoneuria macabaiba sp. nov. (Insecta:
Are the dinosauromorph femora from the Upper Triassic of Hayden Quarry (New Mexico) three stages in a growth series of a single taxon?
 Anais da Academia Brasileira de Ciências (2017) 89(2): 835-839 (Annals of the Brazilian Academy of Sciences) Printed version ISSN 0001-3765 / Online version ISSN 1678-2690 http://dx.doi.org/10.1590/0001-3765201720160583
Anais da Academia Brasileira de Ciências (2017) 89(2): 835-839 (Annals of the Brazilian Academy of Sciences) Printed version ISSN 0001-3765 / Online version ISSN 1678-2690 http://dx.doi.org/10.1590/0001-3765201720160583
O'Regan HJ Defining cheetahs, a multivariante analysis of skull shape in big cats. Mammal Review 32(1):58-62.
 O'Regan HJ. 2002. Defining cheetahs, a multivariante analysis of skull shape in big cats. Mammal Review 32(1):58-62. Keywords: Acinonyx jubatus/cheetah/evolution/felidae/morphology/morphometrics/multivariate
O'Regan HJ. 2002. Defining cheetahs, a multivariante analysis of skull shape in big cats. Mammal Review 32(1):58-62. Keywords: Acinonyx jubatus/cheetah/evolution/felidae/morphology/morphometrics/multivariate
Minnesota_mammals_Info_9.doc 11/04/09 -- DRAFT Page 1 of 64. Minnesota mammals
 Minnesota_mammals_Info_9.doc 11/04/09 -- DRAFT Page 1 of 64 Minnesota mammals This is a short guide to Minnesota mammals, with information drawn from Hazard s Mammals of, Walker s Mammals of the World,
Minnesota_mammals_Info_9.doc 11/04/09 -- DRAFT Page 1 of 64 Minnesota mammals This is a short guide to Minnesota mammals, with information drawn from Hazard s Mammals of, Walker s Mammals of the World,
CHROMOSOMA 9 Springer-Verlag Behaviour of the ZW Sex Bivalent in the Snake Bothrops jararaca. Chromosoma (Berl.) 83, (1981)
 Chromosoma (Berl.) 83, 289-293 (1981) CHROMOSOMA 9 Springer-Verlag 1981 Behaviour of the ZW Sex Bivalent in the Snake Bothrops jararaca Maria Luiza Be~ak* and Willy Be~ak Servigo de Gen~tica, Instituto
Chromosoma (Berl.) 83, 289-293 (1981) CHROMOSOMA 9 Springer-Verlag 1981 Behaviour of the ZW Sex Bivalent in the Snake Bothrops jararaca Maria Luiza Be~ak* and Willy Be~ak Servigo de Gen~tica, Instituto
complex in cusp pattern. (3) The bones of the coyote skull are thinner, crests sharper and the
 DISTINCTIONS BETWEEN THE SKULLS OF S AND DOGS Grover S. Krantz Archaeological sites in the United States frequently yield the bones of coyotes and domestic dogs. These two canines are very similar both
DISTINCTIONS BETWEEN THE SKULLS OF S AND DOGS Grover S. Krantz Archaeological sites in the United States frequently yield the bones of coyotes and domestic dogs. These two canines are very similar both
INTRASPECIFIC AGONISM BETWEEN GIANT OTTER GROUPS. Carolina Ribas 1. Guilherme Mourão 2. Campo Grande, MS , Brazil. Brazil.
 INTRASPECIFIC AGONISM BETWEEN GIANT OTTER GROUPS Carolina Ribas 1 Guilherme Mourão 2 1 Dept. de Biologia- CCBS, Universidade Federal de Mato Grosso do Sul, CP 549, Campo Grande, MS 79070-900, Brazil. 2
INTRASPECIFIC AGONISM BETWEEN GIANT OTTER GROUPS Carolina Ribas 1 Guilherme Mourão 2 1 Dept. de Biologia- CCBS, Universidade Federal de Mato Grosso do Sul, CP 549, Campo Grande, MS 79070-900, Brazil. 2
What are taxonomy, classification, and systematics?
 Topic 2: Comparative Method o Taxonomy, classification, systematics o Importance of phylogenies o A closer look at systematics o Some key concepts o Parts of a cladogram o Groups and characters o Homology
Topic 2: Comparative Method o Taxonomy, classification, systematics o Importance of phylogenies o A closer look at systematics o Some key concepts o Parts of a cladogram o Groups and characters o Homology
Received 16 November 1998; received in revised form and accepted for publication by M. Schmid 8 March 1999
 Chromosome Research 7: 247±254, 1999. # 1999 Kluwer Academic Publishers. Printed in the Netherlands 247 Chromosomal polymorphisms due to supernumerary chromosomes and pericentric inversions in the eyelidless
Chromosome Research 7: 247±254, 1999. # 1999 Kluwer Academic Publishers. Printed in the Netherlands 247 Chromosomal polymorphisms due to supernumerary chromosomes and pericentric inversions in the eyelidless
COMPARATIVE ANALYSIS OF CRANIAL SUTURE COMPLEXITY IN THE GENUS Caiman (CROCODYLIA, ALLIGATORIDAE)
 SUTURE COMPLEXITY IN Caiman 689 COMPARATIVE ANALYSIS OF CRANIAL SUTURE COMPLEXITY IN THE GENUS Caiman (CROCODYLIA, ALLIGATORIDAE) MONTEIRO, L. R. 1 and LESSA, L. G. 2 1 Laboratório de Ciências Ambientais,
SUTURE COMPLEXITY IN Caiman 689 COMPARATIVE ANALYSIS OF CRANIAL SUTURE COMPLEXITY IN THE GENUS Caiman (CROCODYLIA, ALLIGATORIDAE) MONTEIRO, L. R. 1 and LESSA, L. G. 2 1 Laboratório de Ciências Ambientais,
The Karyotype of Plestiodon anthracinus (Baird, 1850) (Sauria: Scincidae): A Step Toward Solving an Enigma
 2017 2017 SOUTHEASTERN Southeastern Naturalist NATURALIST 16(3):326 330 The Karyotype of Plestiodon anthracinus (Baird, 1850) (Sauria: Scincidae): A Step Toward Solving an Enigma Laurence M. Hardy 1, *,
2017 2017 SOUTHEASTERN Southeastern Naturalist NATURALIST 16(3):326 330 The Karyotype of Plestiodon anthracinus (Baird, 1850) (Sauria: Scincidae): A Step Toward Solving an Enigma Laurence M. Hardy 1, *,
A NEW SPECIES OF SCANIA OLIVARES (LEPIDOPTERA, NOCTUIDAE, AUSTRANDESIINI)
 Gayana 69(1): 1-5, 2005 ISSN 0717-652X A NEW SPECIES OF SCANIA OLIVARES (LEPIDOPTERA, NOCTUIDAE, AUSTRANDESIINI) UNA NUEVA ESPECIE DE SCANIA OLIVARES (LEPIDOPTERA, NOCTUIDAE, AUSTRANDESIINI) Tania S. Olivares
Gayana 69(1): 1-5, 2005 ISSN 0717-652X A NEW SPECIES OF SCANIA OLIVARES (LEPIDOPTERA, NOCTUIDAE, AUSTRANDESIINI) UNA NUEVA ESPECIE DE SCANIA OLIVARES (LEPIDOPTERA, NOCTUIDAE, AUSTRANDESIINI) Tania S. Olivares
NOR association in Canis familiaris
 NOR association in Canis familiaris M Rønne, BS Poulsen, Y Shibasaki Odense University, Institute of Medical Biology, Department of Anatomy and Cytology, Campusvej 55, DK-5230 Odense M, Denmark (Proceedings
NOR association in Canis familiaris M Rønne, BS Poulsen, Y Shibasaki Odense University, Institute of Medical Biology, Department of Anatomy and Cytology, Campusvej 55, DK-5230 Odense M, Denmark (Proceedings
New York State Mammals. Order Lagomorpha Order Rodentia
 New York State Mammals Order Lagomorpha Order Rodentia FAMILY: LEPORIDAE Rabbits and hares Conspicuous tail Fenestra appears as bony latticework Some species molt seasonally Presence of a second incisor
New York State Mammals Order Lagomorpha Order Rodentia FAMILY: LEPORIDAE Rabbits and hares Conspicuous tail Fenestra appears as bony latticework Some species molt seasonally Presence of a second incisor
LONG-TAILED FAT-TAILED OPOSSUM Thylamys macrurus (Olfers, 1818)
 LONG-TAILED FAT-TAILED OPOSSUM Thylamys macrurus (Olfers, 1818) FIGURE 1 - Adult, Brazil (Nilton Caceres). TAXONOMY: Class Mammalia; Subclass Theria; Infraclass Metatheria; Magnorder Ameridelphia; Order
LONG-TAILED FAT-TAILED OPOSSUM Thylamys macrurus (Olfers, 1818) FIGURE 1 - Adult, Brazil (Nilton Caceres). TAXONOMY: Class Mammalia; Subclass Theria; Infraclass Metatheria; Magnorder Ameridelphia; Order
OCCASIONAL PAPERS OF THE MUSEUM OF ZOOLOGY UNIVERSITY OF MICHIGAN A NEW PHYLLOTINE RODENT (GENUS GRAOMYS) FROM PARAGUAY
 OCCASIONAL PAPERS OF THE MUSEUM OF ZOOLOGY UNIVERSITY OF MICHIGAN A NEW PHYLLOTINE RODENT (GENUS GRAOMYS) FROM PARAGUAY STUDY OF MAMMALS collected in Paraguay in 1972-73 reveals a new species of the genus
OCCASIONAL PAPERS OF THE MUSEUM OF ZOOLOGY UNIVERSITY OF MICHIGAN A NEW PHYLLOTINE RODENT (GENUS GRAOMYS) FROM PARAGUAY STUDY OF MAMMALS collected in Paraguay in 1972-73 reveals a new species of the genus
Chiroptera Neotropical 14 (1), July First record of Miller's mastiff bat, Molossus pretiosus (Mammalia: Chiroptera), from the Brazilian Caatinga
 First record of Miller's mastiff bat, Molossus pretiosus (Mammalia: Chiroptera), from the Brazilian Caatinga Marcelo R. Nogueira 1*, André Pol 2, Leandro R. Monteiro 3 and Adriano L. Peracchi 2 1. Laboratório
First record of Miller's mastiff bat, Molossus pretiosus (Mammalia: Chiroptera), from the Brazilian Caatinga Marcelo R. Nogueira 1*, André Pol 2, Leandro R. Monteiro 3 and Adriano L. Peracchi 2 1. Laboratório
SUPPLEMENTARY ONLINE MATERIAL FOR. Nirina O. Ratsimbaholison, Ryan N. Felice, and Patrick M. O connor
 http://app.pan.pl/som/app61-ratsimbaholison_etal_som.pdf SUPPLEMENTARY ONLINE MATERIAL FOR Nirina O. Ratsimbaholison, Ryan N. Felice, and Patrick M. O connor Ontogenetic changes in the craniomandibular
http://app.pan.pl/som/app61-ratsimbaholison_etal_som.pdf SUPPLEMENTARY ONLINE MATERIAL FOR Nirina O. Ratsimbaholison, Ryan N. Felice, and Patrick M. O connor Ontogenetic changes in the craniomandibular
New species of Cerradomys from coastal sandy plains of southeastern Brazil (Cricetidae: Sigmodontinae)
 Journal of Mammalogy, 92(3):645 658, 2011 New species of Cerradomys from coastal sandy plains of southeastern Brazil (Cricetidae: Sigmodontinae) WILLIAM CORRÊA TAVARES, LEILA MARIA PESSÔA,* AND PABLO RODRIGUES
Journal of Mammalogy, 92(3):645 658, 2011 New species of Cerradomys from coastal sandy plains of southeastern Brazil (Cricetidae: Sigmodontinae) WILLIAM CORRÊA TAVARES, LEILA MARIA PESSÔA,* AND PABLO RODRIGUES
Three new species of Microctenochira SPAETH from Brazil and Panama (Coleoptera: Chrysomelidae: Cassidinae)
 Genus Vol. 10 (1): 109-116 Wroc³aw, 31 III 1999 Three new species of Microctenochira SPAETH from Brazil and Panama (Coleoptera: Chrysomelidae: Cassidinae) JOLANTA ŒWIÊTOJAÑSKA and LECH BOROWIEC Zoological
Genus Vol. 10 (1): 109-116 Wroc³aw, 31 III 1999 Three new species of Microctenochira SPAETH from Brazil and Panama (Coleoptera: Chrysomelidae: Cassidinae) JOLANTA ŒWIÊTOJAÑSKA and LECH BOROWIEC Zoological
Corresponding author: M.C. Barros
 Phylogeny of Marmosops and the occurrence of Marmosops pinheiroi (Pine, 1981) (Didelphimorphia, Didelphidae) in the Cerrado savanna of Maranhão, Brazil D.C. Nascimento 1,2, A.P.M. Olímpio 2, E. Conceição
Phylogeny of Marmosops and the occurrence of Marmosops pinheiroi (Pine, 1981) (Didelphimorphia, Didelphidae) in the Cerrado savanna of Maranhão, Brazil D.C. Nascimento 1,2, A.P.M. Olímpio 2, E. Conceição
A new species of the genus Phytocoris (Heteroptera: Miridae) from the United Arab Emirates
 ACTA ENTOMOLOGICA MUSEI NATIONALIS PRAGAE Published 6.xi.2006 Volume 46, pp. 15-19 ISSN 0374-1036 A new species of the genus Phytocoris (Heteroptera: Miridae) from the United Arab Emirates Rauno E. LINNAVUORI
ACTA ENTOMOLOGICA MUSEI NATIONALIS PRAGAE Published 6.xi.2006 Volume 46, pp. 15-19 ISSN 0374-1036 A new species of the genus Phytocoris (Heteroptera: Miridae) from the United Arab Emirates Rauno E. LINNAVUORI
FURTHER STUDIES ON TWO SKELETONS OF THE BLACK RIGHT WHALE IN THE NORTH PACIFIC
 FURTHER STUDIES ON TWO SKELETONS OF THE BLACK RIGHT WHALE IN THE NORTH PACIFIC HIDEO OMURA, MASAHARU NISHIWAKI* AND TOSHIO KASUYA* ABSTRACT Two skeletons of the black right whale were studied, supplementing
FURTHER STUDIES ON TWO SKELETONS OF THE BLACK RIGHT WHALE IN THE NORTH PACIFIC HIDEO OMURA, MASAHARU NISHIWAKI* AND TOSHIO KASUYA* ABSTRACT Two skeletons of the black right whale were studied, supplementing
CENE RUMINANTS OF THE GENERA OVIBOS AND
 DESCRIPTIONS OF TWO NEW SPECIES OF PLEISTO- CENE RUMINANTS OF THE GENERA OVIBOS AND BOOTHERIUM, WITH NOTES ON THE LATTER GENUS. By James Williams Gidley, Of the United States National Museum. Two interesting
DESCRIPTIONS OF TWO NEW SPECIES OF PLEISTO- CENE RUMINANTS OF THE GENERA OVIBOS AND BOOTHERIUM, WITH NOTES ON THE LATTER GENUS. By James Williams Gidley, Of the United States National Museum. Two interesting
Sergio, A NEW GENUS OF GHOST SHRIMP FROM THE AMERICAS (CRUSTACEA: DECAPODA: CALLIANASSIDAE)
 NAUPLIUS, Rio Grande, 1: 39-43, 1991!* ^ Sergio, A NEW GENUS OF GHOST SHRIMP FROM THE AMERICAS (CRUSTACEA: DECAPODA: CALLIANASSIDAE) R. B. MANNING & R. LEMAITRE Department of Invertebrate Zoology National
NAUPLIUS, Rio Grande, 1: 39-43, 1991!* ^ Sergio, A NEW GENUS OF GHOST SHRIMP FROM THE AMERICAS (CRUSTACEA: DECAPODA: CALLIANASSIDAE) R. B. MANNING & R. LEMAITRE Department of Invertebrate Zoology National
Wild Fur Identification. an identification aid for Lynx species fur
 Wild Fur Identification an identification aid for Lynx species fur Wild Fur Identifica- -an identification and classification aid for Lynx species fur pelts. Purpose: There are four species of Lynx including
Wild Fur Identification an identification aid for Lynx species fur Wild Fur Identifica- -an identification and classification aid for Lynx species fur pelts. Purpose: There are four species of Lynx including
Diagnosis of Living and Fossil Short-necked Turtles of the Genus Elseya using skeletal morphology
 Diagnosis of Living and Fossil Short-necked Turtles of the Genus Elseya using skeletal morphology by Scott Andrew Thomson B.App.Sc. University of Canberra Institute of Applied Ecology University of Canberra
Diagnosis of Living and Fossil Short-necked Turtles of the Genus Elseya using skeletal morphology by Scott Andrew Thomson B.App.Sc. University of Canberra Institute of Applied Ecology University of Canberra
Central Marine Fisheries Research Institute, Mandapam Camp
 w«r n Mar. biol. Ass. India, 1961, 3 (1 & 2): 92-95 ON A NEW GENUS OF PORCELLANIDAE (CRUSTACEA-ANOMURA) * By C. SANKARANKUTTY Central Marine Fisheries Research Institute, Mandapam Camp The specimen described
w«r n Mar. biol. Ass. India, 1961, 3 (1 & 2): 92-95 ON A NEW GENUS OF PORCELLANIDAE (CRUSTACEA-ANOMURA) * By C. SANKARANKUTTY Central Marine Fisheries Research Institute, Mandapam Camp The specimen described
Diagnosis of Leptospira spp. Infection in Sheep Flocks in the State of Mato Grosso, Brazil
 Acta Scientiae Veterinariae, 2017. 45: 1499. RESEARCH ARTICLE Pub. 1499 ISSN 1679-9216 Diagnosis of Leptospira spp. Infection in Sheep Flocks in the State of Mato Grosso, Brazil Camila Eckstein 1, Luciano
Acta Scientiae Veterinariae, 2017. 45: 1499. RESEARCH ARTICLE Pub. 1499 ISSN 1679-9216 Diagnosis of Leptospira spp. Infection in Sheep Flocks in the State of Mato Grosso, Brazil Camila Eckstein 1, Luciano
SERGIO SOLARI 1, 3 & RONALD H. PINE 2. Abstract. Resumen
 Zootaxa 1756: 49 61 (2008) www.mapress.com/zootaxa/ Copyright 2008 Magnolia Press ISSN 1175-5326 (print edition) ZOOTAXA ISSN 1175-5334 (online edition) Rediscovery and redescription of Marmosa (Stegomarmosa)
Zootaxa 1756: 49 61 (2008) www.mapress.com/zootaxa/ Copyright 2008 Magnolia Press ISSN 1175-5326 (print edition) ZOOTAXA ISSN 1175-5334 (online edition) Rediscovery and redescription of Marmosa (Stegomarmosa)
107. Segregation o f Karyotypes in the F2 Generation o f the Hybrids between Mauritius and Oceanian Type Black Rats with a Note on their Litter Size*'
 No. 9] Proc. Japan Acad., 5'6, Ser. B (1980) 557 107. Segregation o f Karyotypes in the F2 Generation o f the Hybrids between Mauritius and Oceanian Type Black Rats with a Note on their Litter Size*' By
No. 9] Proc. Japan Acad., 5'6, Ser. B (1980) 557 107. Segregation o f Karyotypes in the F2 Generation o f the Hybrids between Mauritius and Oceanian Type Black Rats with a Note on their Litter Size*' By
New York State Mammals. Morphology Ecology Identification Classification Distribution
 New York State Mammals Morphology Ecology Identification Classification Distribution ORDER: Didelphimorphia FAMILY: Didelphidae Common Name: Virginia opossum Scientific Name: (Didelphis virginiana) Marsupial
New York State Mammals Morphology Ecology Identification Classification Distribution ORDER: Didelphimorphia FAMILY: Didelphidae Common Name: Virginia opossum Scientific Name: (Didelphis virginiana) Marsupial
PARTIAL REPORT. Juvenile hybrid turtles along the Brazilian coast RIO GRANDE FEDERAL UNIVERSITY
 RIO GRANDE FEDERAL UNIVERSITY OCEANOGRAPHY INSTITUTE MARINE MOLECULAR ECOLOGY LABORATORY PARTIAL REPORT Juvenile hybrid turtles along the Brazilian coast PROJECT LEADER: MAIRA PROIETTI PROFESSOR, OCEANOGRAPHY
RIO GRANDE FEDERAL UNIVERSITY OCEANOGRAPHY INSTITUTE MARINE MOLECULAR ECOLOGY LABORATORY PARTIAL REPORT Juvenile hybrid turtles along the Brazilian coast PROJECT LEADER: MAIRA PROIETTI PROFESSOR, OCEANOGRAPHY
Aging by molt patterns of flight feathers of non adult Steller s Sea Eagle
 First Symposium on Steller s and White-tailed Sea Eagles in East Asia pp. 11-16, 2000 UETA, M. & MCGRADY, M.J. (eds) Wild Bird Society of Japan, Tokyo Japan Aging by molt patterns of flight feathers of
First Symposium on Steller s and White-tailed Sea Eagles in East Asia pp. 11-16, 2000 UETA, M. & MCGRADY, M.J. (eds) Wild Bird Society of Japan, Tokyo Japan Aging by molt patterns of flight feathers of
AMERICAN MUSEUM NOVITATES Published by
 AMERICAN MUSEUM NOVITATES Published by Number 782 THE AmzRICAN MUSEUM OF NATURAL HISTORY Feb. 20, 1935 New York City 56.81, 7 G (68) A NOTE ON THE CYNODONT, GLOCHINODONTOIDES GRACILIS HAUGHTON BY LIEUWE
AMERICAN MUSEUM NOVITATES Published by Number 782 THE AmzRICAN MUSEUM OF NATURAL HISTORY Feb. 20, 1935 New York City 56.81, 7 G (68) A NOTE ON THE CYNODONT, GLOCHINODONTOIDES GRACILIS HAUGHTON BY LIEUWE
Article urn:lsid:zoobank.org:pub:c7c52eea-b748-4f05-813d-92acf74821a3
 7 Zootaxa 3481: 60 72 (2012) www.mapress.com/zootaxa/ Copyright 2012 Magnolia Press Article urn:lsid:zoobank.org:pub:c7c52eea-b748-4f05-813d-92acf74821a3 ISSN 1175-5326 (print edition) ZOOTAXA ISSN 1175-5334
7 Zootaxa 3481: 60 72 (2012) www.mapress.com/zootaxa/ Copyright 2012 Magnolia Press Article urn:lsid:zoobank.org:pub:c7c52eea-b748-4f05-813d-92acf74821a3 ISSN 1175-5326 (print edition) ZOOTAXA ISSN 1175-5334
Minnesota_mammals_Info_12.doc 11/20/09 -- DRAFT Page 36 of 42
 Minnesota_mammals_Info_12.doc 11/20/09 -- DRAFT Page 36 of 42 The Families Muridae and Cricetidae. As we discussed in class, these familes are now separated again. At one point the Muridae included cricetids
Minnesota_mammals_Info_12.doc 11/20/09 -- DRAFT Page 36 of 42 The Families Muridae and Cricetidae. As we discussed in class, these familes are now separated again. At one point the Muridae included cricetids
Appendix 4: Keys to the bats of the Greater Yellowstone Network
 Appendix 4: Keys to the bats of the Greater Yellowstone Network Page 66 Dichotomous Key to the Bats of the Greater Yellowstone Network Doug Keinath, WYNDD, dkeinath@uwyo.edu # If this is true then go to
Appendix 4: Keys to the bats of the Greater Yellowstone Network Page 66 Dichotomous Key to the Bats of the Greater Yellowstone Network Doug Keinath, WYNDD, dkeinath@uwyo.edu # If this is true then go to
Polecats & Ferrets. How to tell them apart
 Polecats & Ferrets How to tell them apart Introduction The polecat (Mustela putorius) is expanding its range in Britain, and in many areas across Britain, ferrets (Mustela furo) occur either as individuals
Polecats & Ferrets How to tell them apart Introduction The polecat (Mustela putorius) is expanding its range in Britain, and in many areas across Britain, ferrets (Mustela furo) occur either as individuals
W. R. Heyer, 1 R. O. de Sá, 2 and A. Rettig 2. Herpetologia Petropolitana, Ananjeva N. and Tsinenko O. (eds.), pp
 Herpetologia Petropolitana, Ananjeva N. and Tsinenko O. (eds.), pp. 35 39 35 SIBLING SPECIES, ADVERTISEMENT CALLS, AND REPRODUCTIVE ISOLATION IN FROGS OF THE Leptodactylus pentadactylus SPECIES CLUSTER
Herpetologia Petropolitana, Ananjeva N. and Tsinenko O. (eds.), pp. 35 39 35 SIBLING SPECIES, ADVERTISEMENT CALLS, AND REPRODUCTIVE ISOLATION IN FROGS OF THE Leptodactylus pentadactylus SPECIES CLUSTER
Lab 5: Rodentia and Lagomorpha
 Lab 5: Rodentia and Lagomorpha (8 families in B.C.) Sciuridae squirrels (16 species in B.C.) Muridae mice, rats, lemmings, voles (16) Aplodontidae mountain beaver (1) Castoridae beaver (1) Dipodidae jumping
Lab 5: Rodentia and Lagomorpha (8 families in B.C.) Sciuridae squirrels (16 species in B.C.) Muridae mice, rats, lemmings, voles (16) Aplodontidae mountain beaver (1) Castoridae beaver (1) Dipodidae jumping
Two new Phradonoma species (Coleoptera: Dermestidae) from Iran
 Journal of Entomological Society of Iran 2008, 28(1), 87-91 87 Two new Phradonoma species (Coleoptera: Dermestidae) from Iran A. Herrmann 1&* and J. Háva 2 1. Bremervörder Strasse 123, D - 21682 Stade,
Journal of Entomological Society of Iran 2008, 28(1), 87-91 87 Two new Phradonoma species (Coleoptera: Dermestidae) from Iran A. Herrmann 1&* and J. Háva 2 1. Bremervörder Strasse 123, D - 21682 Stade,
DNA-based and geometric morphometric analysis to validate species designation: a case study of the subterranean rodent Ctenomys bicolor
 DNA-based and geometric morphometric analysis to validate species designation: a case study of the subterranean rodent Ctenomys bicolor J.F.B. Stolz, G.L. Gonçalves, L. Leipnitz and T.R.O. Freitas Laboratório
DNA-based and geometric morphometric analysis to validate species designation: a case study of the subterranean rodent Ctenomys bicolor J.F.B. Stolz, G.L. Gonçalves, L. Leipnitz and T.R.O. Freitas Laboratório
SUBFAMILY THYMOPINAE Holthuis, 1974
 click for previous page 29 Remarks : The taxonomy of the species is not clear. It is possible that 2 forms may have to be distinguished: A. sublevis Wood-Mason, 1891 (with a synonym A. opipara Burukovsky
click for previous page 29 Remarks : The taxonomy of the species is not clear. It is possible that 2 forms may have to be distinguished: A. sublevis Wood-Mason, 1891 (with a synonym A. opipara Burukovsky
ZOOLOGISCHE MEDEDELINGEN
 MINISTERIE VAN ONDERWIJS, KUNSTEN EN WETENSCHAPPEN ZOOLOGISCHE MEDEDELINGEN UITGEGEVEN DOOR HET RIJKSMUSEUM VAN NATUURLIJKE HISTORIE TE LEIDEN DEEL XXXIII, No. 10 13 December 1954 ON VAMPYRODES CARACCIOLAE
MINISTERIE VAN ONDERWIJS, KUNSTEN EN WETENSCHAPPEN ZOOLOGISCHE MEDEDELINGEN UITGEGEVEN DOOR HET RIJKSMUSEUM VAN NATUURLIJKE HISTORIE TE LEIDEN DEEL XXXIII, No. 10 13 December 1954 ON VAMPYRODES CARACCIOLAE
BULLETIN OF THE ALLYN MUSEUM
 BULLETIN OF THE ALLYN MUSEUM 3701 Bayshore Rd. Sarasota, Florida 33580 Number 100 Published By The Florida State Museum University of Florida Gainesville. Florida 32611 30 January 1986 A PRELIMINARY REVISION
BULLETIN OF THE ALLYN MUSEUM 3701 Bayshore Rd. Sarasota, Florida 33580 Number 100 Published By The Florida State Museum University of Florida Gainesville. Florida 32611 30 January 1986 A PRELIMINARY REVISION
v:ii-ixi, 'i':;iisimvi'\>!i-:: "^ A%'''''-'^-''S.''v.--..V^'E^'-'-^"-t''gi L I E) R.ARY OF THE VERSITY U N I or ILLINOIS REMO
 "^ A%'''''-'^-''S.''v.--..V^'E^'-'-^"-t''gi v:ii-ixi, 'i':;iisimvi'\>!i-:: L I E) R.ARY OF THE U N I VERSITY or ILLINOIS REMO Natural History Survey Librarv GEOLOGICAL SERIES OF FIELD MUSEUM OF NATURAL
"^ A%'''''-'^-''S.''v.--..V^'E^'-'-^"-t''gi v:ii-ixi, 'i':;iisimvi'\>!i-:: L I E) R.ARY OF THE U N I VERSITY or ILLINOIS REMO Natural History Survey Librarv GEOLOGICAL SERIES OF FIELD MUSEUM OF NATURAL
New York State Mammals. Order Rodentia (cont.) Order Lagomorpha
 New York State Mammals Order Rodentia (cont.) Order Lagomorpha FAMILY: CRICETIDAE New World rats, mice, voles, hamsters, etc. Diverse & species rich Most terrestrial, 1 in NYS is aquatic Muskrat Subfamily
New York State Mammals Order Rodentia (cont.) Order Lagomorpha FAMILY: CRICETIDAE New World rats, mice, voles, hamsters, etc. Diverse & species rich Most terrestrial, 1 in NYS is aquatic Muskrat Subfamily
Cytogenetic Study of the Leopard, Panthera pardus (Carnivora, Felidae) by Conventional Staining, G- banding and High-resolution Staining Technique
 2008 The Japan Mendel Society Cytologia 73(1): 81 90, 2008 Cytogenetic Study of the Leopard, Panthera pardus (Carnivora, Felidae) by Conventional Staining, G- banding and High-resolution Staining Technique
2008 The Japan Mendel Society Cytologia 73(1): 81 90, 2008 Cytogenetic Study of the Leopard, Panthera pardus (Carnivora, Felidae) by Conventional Staining, G- banding and High-resolution Staining Technique
Effect of Cage Density on the Performance of 25- to 84-Week-Old Laying Hens
 Brazilian Journal of Poultry Science Revista Brasileira de Ciência Avícola ISSN 1516-635X Oct - Dec 2009 / v.11 / n.4 / 257-262 Effect of Cage Density on the Performance of 25- to 84- Author(s) Rios RL
Brazilian Journal of Poultry Science Revista Brasileira de Ciência Avícola ISSN 1516-635X Oct - Dec 2009 / v.11 / n.4 / 257-262 Effect of Cage Density on the Performance of 25- to 84- Author(s) Rios RL
Williston, and as there are many fairly good specimens in the American
 56.81.7D :14.71.5 Article VII.- SOME POINTS IN THE STRUCTURE OF THE DIADECTID SKULL. BY R. BROOM. The skull of Diadectes has been described by Cope, Case, v. Huene, and Williston, and as there are many
56.81.7D :14.71.5 Article VII.- SOME POINTS IN THE STRUCTURE OF THE DIADECTID SKULL. BY R. BROOM. The skull of Diadectes has been described by Cope, Case, v. Huene, and Williston, and as there are many
Taxonomic status and relationships of Sorex obscurus parvidens Jackson, 1921, from California
 Journal of Mammalogy, 93(3):826 838, 2012 Taxonomic status and relationships of Sorex obscurus parvidens Jackson, 1921, from California NEAL WOODMAN* United States Geological Survey Patuxent Wildlife Research
Journal of Mammalogy, 93(3):826 838, 2012 Taxonomic status and relationships of Sorex obscurus parvidens Jackson, 1921, from California NEAL WOODMAN* United States Geological Survey Patuxent Wildlife Research
Mastozoología Neotropical ISSN: Sociedad Argentina para el Estudio de los Mamíferos. Argentina
 Mastozoología Neotropical ISSN: 0327-9383 ulyses@cenpat.edu.ar Sociedad Argentina para el Estudio de los Mamíferos Argentina Díaz-N., Juan F.; Gómez-Laverde, Marcela; Sánchez-Giraldo, Camilo Rediscovery
Mastozoología Neotropical ISSN: 0327-9383 ulyses@cenpat.edu.ar Sociedad Argentina para el Estudio de los Mamíferos Argentina Díaz-N., Juan F.; Gómez-Laverde, Marcela; Sánchez-Giraldo, Camilo Rediscovery
A new species of Amphisbaena (Squamata, Amphisbaenidae) from state of Maranhão, Brazil
 A new species of Amphisbaena (Squamata, Amphisbaenidae) from state of Maranhão, Brazil Miguel Trefaut Rodrigues 1, Gilda V. Andrade 2 and Jucivaldo Dias Lima 2 Phyllomedusa 2(1):21-26, 2003 2003 Melopsittacus
A new species of Amphisbaena (Squamata, Amphisbaenidae) from state of Maranhão, Brazil Miguel Trefaut Rodrigues 1, Gilda V. Andrade 2 and Jucivaldo Dias Lima 2 Phyllomedusa 2(1):21-26, 2003 2003 Melopsittacus
2007 The Japan Mendel Society Cytologia 72(1): , 2007
 2007 The Japan Mendel Society Cytologia 72(1): 101 110, 2007 A Study on Karyotype of the Asian Leopard Cat, Prionailurus bengalensis (Carnivora, Felidae) by Conventional Staining, G-banding and High-resolution
2007 The Japan Mendel Society Cytologia 72(1): 101 110, 2007 A Study on Karyotype of the Asian Leopard Cat, Prionailurus bengalensis (Carnivora, Felidae) by Conventional Staining, G-banding and High-resolution
posterior part of the second segment may show a few white hairs
 April, 1911.] New Species of Diptera of the Genus Erax. 307 NEW SPECIES OF DIPTERA OF THE GENUS ERAX. JAMES S. HINE. The various species of Asilinae known by the generic name Erax have been considered
April, 1911.] New Species of Diptera of the Genus Erax. 307 NEW SPECIES OF DIPTERA OF THE GENUS ERAX. JAMES S. HINE. The various species of Asilinae known by the generic name Erax have been considered
A NEW SPECIES OF A USTROLIBINIA FROM THE SOUTH CHINA SEA AND INDONESIA (CRUSTACEA: BRACHYURA: MAJIDAE)
 69 C O a g r ^ j^a RAFFLES BULLETIN OF ZOOLOGY 1992 40(1): 69-73 A NEW SPECIES OF A USTROLIBINIA FROM THE SOUTH CHINA SEA AND INDONESIA (CRUSTACEA: BRACHYURA: MAJIDAE) H P Waener SMITHSONIAN INSTITUTE
69 C O a g r ^ j^a RAFFLES BULLETIN OF ZOOLOGY 1992 40(1): 69-73 A NEW SPECIES OF A USTROLIBINIA FROM THE SOUTH CHINA SEA AND INDONESIA (CRUSTACEA: BRACHYURA: MAJIDAE) H P Waener SMITHSONIAN INSTITUTE
HONR219D Due 3/29/16 Homework VI
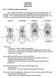 Part 1: Yet More Vertebrate Anatomy!!! HONR219D Due 3/29/16 Homework VI Part 1 builds on homework V by examining the skull in even greater detail. We start with the some of the important bones (thankfully
Part 1: Yet More Vertebrate Anatomy!!! HONR219D Due 3/29/16 Homework VI Part 1 builds on homework V by examining the skull in even greater detail. We start with the some of the important bones (thankfully
SENSITIZATION FOR THE AUTOCHTHONOUS BREEDS CONSERVATION VIA THE PUBLIC SHOWS OF ANIMALS
 SENSITIZATION FOR THE AUTOCHTHONOUS BREEDS CONSERVATION VIA THE PUBLIC SHOWS OF ANIMALS SENSIBILIZACION DE LA OPINION PUBLICA POR LA CONSERVACION DE RAZAS AUTOCTONAS A TRAVES DE LAS EXPOSICIONES DE ANIMALES
SENSITIZATION FOR THE AUTOCHTHONOUS BREEDS CONSERVATION VIA THE PUBLIC SHOWS OF ANIMALS SENSIBILIZACION DE LA OPINION PUBLICA POR LA CONSERVACION DE RAZAS AUTOCTONAS A TRAVES DE LAS EXPOSICIONES DE ANIMALES
ALFRED L. GARDNER 1 AND MICHAEL D. CARLETON 2 ABSTRACT
 Chapter 5 A New Species of Reithrodontomys, Subgenus Aporodon (Cricetidae: Neotominae), from the Highlands of Costa Rica, with Comments on Costa Rican and Panamanian Reithrodontomys ALFRED L. GARDNER 1
Chapter 5 A New Species of Reithrodontomys, Subgenus Aporodon (Cricetidae: Neotominae), from the Highlands of Costa Rica, with Comments on Costa Rican and Panamanian Reithrodontomys ALFRED L. GARDNER 1
DISTRIBUTION AND MORPHOMETRICS OF NATALUS STRAMINEUS FROM SOUTH AMERICA (CHIROPTERA, NATALIDAE)
 Distribution and morphometrics of Natalus stramineus from South America... 123 DISTRIBUTION AND MORPHOMETRICS OF NATALUS STRAMINEUS FROM SOUTH AMERICA (CHIROPTERA, NATALIDAE) ABSTRACT Valdir Antonio Taddei
Distribution and morphometrics of Natalus stramineus from South America... 123 DISTRIBUTION AND MORPHOMETRICS OF NATALUS STRAMINEUS FROM SOUTH AMERICA (CHIROPTERA, NATALIDAE) ABSTRACT Valdir Antonio Taddei
First record of visual displays in Scinax cardosoi (Anura: Hylidae)
 Short CommuniCation First record of visual displays in Scinax cardosoi (Anura: Hylidae) Matheus de Toledo Moroti, 1 Mariana Pedrozo, 2 Guilherme Sestito, 1 and Diego José Santana 1 1 970, Campo Grande,
Short CommuniCation First record of visual displays in Scinax cardosoi (Anura: Hylidae) Matheus de Toledo Moroti, 1 Mariana Pedrozo, 2 Guilherme Sestito, 1 and Diego José Santana 1 1 970, Campo Grande,
Marsupial Mole. Notoryctes species. Amy Mutton Zoologist Species and Communities Branch Science and Conservation Division
 Marsupial Mole Notoryctes species Amy Mutton Zoologist Species and Communities Branch Science and Conservation Division Scientific classification Kingdom: Phylum: Class: Infraclass: Order: Family: Animalia
Marsupial Mole Notoryctes species Amy Mutton Zoologist Species and Communities Branch Science and Conservation Division Scientific classification Kingdom: Phylum: Class: Infraclass: Order: Family: Animalia
Mammalogy Laboratory 1 - Mammalian Anatomy
 Mammalogy Laboratory 1 - Mammalian Anatomy I. The Goal. The goal of the lab is to teach you skeletal anatomy of mammals. We will emphasize the skull because many of the taxonomically important characters
Mammalogy Laboratory 1 - Mammalian Anatomy I. The Goal. The goal of the lab is to teach you skeletal anatomy of mammals. We will emphasize the skull because many of the taxonomically important characters
Supplementary Information for: 3D morphometric analysis of fossil canid skulls contradicts
 Supplementary Information for: 3D morphometric analysis of fossil canid skulls contradicts the suggested domestication of dogs during the late Paleolithic Abby Grace Drake 1, * Michael Coquerelle 2,3 Guillaume
Supplementary Information for: 3D morphometric analysis of fossil canid skulls contradicts the suggested domestication of dogs during the late Paleolithic Abby Grace Drake 1, * Michael Coquerelle 2,3 Guillaume
AMERICAN MUSEUM NOVITATES Publiished by
 AMERICAN MUSEUM NOVITATES Publiished by Number 802 THU AmERICAN Mueum of NATURAL HISTORY May 18, 1935 New York City 59.9, 32 R (9) RESULTS OF THE ARCHBOLD EXPEDITIONS. NO. 2 TWELVE APPARENTLY NEW FORMS
AMERICAN MUSEUM NOVITATES Publiished by Number 802 THU AmERICAN Mueum of NATURAL HISTORY May 18, 1935 New York City 59.9, 32 R (9) RESULTS OF THE ARCHBOLD EXPEDITIONS. NO. 2 TWELVE APPARENTLY NEW FORMS
Ciasg\ \;"^iaj?te_. --^::^^5f5c
 Ciasg\ \;"^iaj?te_ --^::^^5f5c NEW PERUVIAN MAMMALS BY WILFRED H. OSGOOD. The mammals described below are those obviously new from a collection made during the past year in northern Peru by Mr. M.
Ciasg\ \;"^iaj?te_ --^::^^5f5c NEW PERUVIAN MAMMALS BY WILFRED H. OSGOOD. The mammals described below are those obviously new from a collection made during the past year in northern Peru by Mr. M.
occasional papers, museum of texas tech university
 Occasional Papers Museum of Texas Tech University Number 326 26 June 2014 New Species of Scotophilus (Chiroptera: Vespertilionidae) from Sub-Saharan Africa Daniel M. Brooks and John W. Bickham Abstract
Occasional Papers Museum of Texas Tech University Number 326 26 June 2014 New Species of Scotophilus (Chiroptera: Vespertilionidae) from Sub-Saharan Africa Daniel M. Brooks and John W. Bickham Abstract
oxfitates Mllsdum M ie'ican Group of Lizards in the Genus Sceloporusl Karyotypes and Evolution of the spinosus COLE2 BY CHARLES J.
 M ie'ican Mllsdum oxfitates PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, N. Y. I0024 NUMBER 243I SEPTEMBER 28, 1970 Karyotypes and Evolution of the spinosus
M ie'ican Mllsdum oxfitates PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, N. Y. I0024 NUMBER 243I SEPTEMBER 28, 1970 Karyotypes and Evolution of the spinosus
Gulf and Caribbean Research
 Gulf and Caribbean Research Volume 16 Issue 1 January 4 Morphological Characteristics of the Carapace of the Hawksbill Turtle, Eretmochelys imbricata, from n Waters Mari Kobayashi Hokkaido University DOI:
Gulf and Caribbean Research Volume 16 Issue 1 January 4 Morphological Characteristics of the Carapace of the Hawksbill Turtle, Eretmochelys imbricata, from n Waters Mari Kobayashi Hokkaido University DOI:
By H. G. JOHNSTON, Ames, Iowa.
 Dec., 19930 Bulletin of the Brooklyn Entomological Society 295 FOUR NEW SPECIES OF MIRIDAE FROM TEXAS (HEMIPTERA).* By H. G. JOHNSTON, Ames, Iowa. Phytocoris conspicuus n. sp. This species is readily distinguished
Dec., 19930 Bulletin of the Brooklyn Entomological Society 295 FOUR NEW SPECIES OF MIRIDAE FROM TEXAS (HEMIPTERA).* By H. G. JOHNSTON, Ames, Iowa. Phytocoris conspicuus n. sp. This species is readily distinguished
Revista Chilena de Historia Natural ISSN: X Sociedad de Biología de Chile Chile
 Revista Chilena de Historia Natural ISSN: 0716-078X editorial@revchilhistnat.com Sociedad de Biología de Chile Chile DE SOUZA, ANDRÉ R.; SILVA, NEWTON J. J.; PREZOTO, FÁBIO A rare but successful reproductive
Revista Chilena de Historia Natural ISSN: 0716-078X editorial@revchilhistnat.com Sociedad de Biología de Chile Chile DE SOUZA, ANDRÉ R.; SILVA, NEWTON J. J.; PREZOTO, FÁBIO A rare but successful reproductive
Article. Abstract ANDRES PARADA 1, AGUSTINA OJEDA 2, SOLANA TABENI 2 & GUILLERMO D'ELÍA 3 1. Introduction
 Zootaxa 3402: 61 68 (2012) www.mapress.com/zootaxa/ Copyright 2012 Magnolia Press Article ISSN 1175-5326 (print edition) ZOOTAXA ISSN 1175-5334 (online edition) The population of Ctenomys from the Ñacuñán
Zootaxa 3402: 61 68 (2012) www.mapress.com/zootaxa/ Copyright 2012 Magnolia Press Article ISSN 1175-5326 (print edition) ZOOTAXA ISSN 1175-5334 (online edition) The population of Ctenomys from the Ñacuñán
SCIUROPTERUS MINDANENSIS SP. NOV., A NEW SPECIES OF FLYING SQUIRREL FROM MINDANAO
 SCIUROPTERUS MINDANENSIS SP. NOV., A NEW SPECIES OF FLYING SQUIRREL FROM MINDANAO By DioscoRO S. Rabor Of the Division of Fisheries^ Department of Agriculture and Commerce Manila FOUR PLATES In August,
SCIUROPTERUS MINDANENSIS SP. NOV., A NEW SPECIES OF FLYING SQUIRREL FROM MINDANAO By DioscoRO S. Rabor Of the Division of Fisheries^ Department of Agriculture and Commerce Manila FOUR PLATES In August,
A Revision of Extant Greater Antillean Bats of the Genus Natalus
 PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, NY 10024 Number 3493, 22 pp., 7 figures, 2 tables October 27, 2005 A Revision of Extant Greater Antillean
PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, NY 10024 Number 3493, 22 pp., 7 figures, 2 tables October 27, 2005 A Revision of Extant Greater Antillean
THE SKULLS OF ARAEOSCELIS AND CASEA, PERMIAN REPTILES
 THE SKULLS OF REOSCELIS ND CSE, PERMIN REPTILES University of Chicago There are few Permian reptiles of greater interest at the present time than the peculiar one I briefly described in this journal' three
THE SKULLS OF REOSCELIS ND CSE, PERMIN REPTILES University of Chicago There are few Permian reptiles of greater interest at the present time than the peculiar one I briefly described in this journal' three
Morphology and geographical distribution of the poorly known snake Umbrivaga pygmaea (Serpentes: Dipsadidae) in Brazil
 Phyllomedusa 10(2):177 182, 2011 2011 Departamento de Ciências Biológicas - ESALQ - USP ISSN 1519-1397 Short Communication Morphology and geographical distribution of the poorly known snake Umbrivaga pygmaea
Phyllomedusa 10(2):177 182, 2011 2011 Departamento de Ciências Biológicas - ESALQ - USP ISSN 1519-1397 Short Communication Morphology and geographical distribution of the poorly known snake Umbrivaga pygmaea
ZOOLOGISCHE MEDEDELINGEN
 ZOOLOGISCHE MEDEDELINGEN U I T G E G E V E N D O O R H E T RIJKSMUSEUM VAN NATUURLIJKE HISTORIE TE LEIDEN (MINISTERIE V A N CULTUUR, RECREATIE E N MAATSCHAPPELIJK WERK) Deel 45 no. 13 15 maart 1971 THE
ZOOLOGISCHE MEDEDELINGEN U I T G E G E V E N D O O R H E T RIJKSMUSEUM VAN NATUURLIJKE HISTORIE TE LEIDEN (MINISTERIE V A N CULTUUR, RECREATIE E N MAATSCHAPPELIJK WERK) Deel 45 no. 13 15 maart 1971 THE
Soleglad, Fet & Lowe: Hadrurus spadix Subgroup
 9 Figures 3 17: Carapace pattern schemes for the Hadrurus arizonensis group. 3. H. arizonensis arizonensis, juvenile male, typical dark phenotype, Rte 178, 0.5 W Rte 127, Inyo Co., California, USA. 4.
9 Figures 3 17: Carapace pattern schemes for the Hadrurus arizonensis group. 3. H. arizonensis arizonensis, juvenile male, typical dark phenotype, Rte 178, 0.5 W Rte 127, Inyo Co., California, USA. 4.
ANTHR 1L Biological Anthropology Lab
 ANTHR 1L Biological Anthropology Lab Name: DEFINING THE ORDER PRIMATES Humans belong to the zoological Order Primates, which is one of the 18 Orders of the Class Mammalia. Today we will review some of
ANTHR 1L Biological Anthropology Lab Name: DEFINING THE ORDER PRIMATES Humans belong to the zoological Order Primates, which is one of the 18 Orders of the Class Mammalia. Today we will review some of
Article.
 Zootaxa 3753 (2): 196 200 www.mapress.com/zootaxa/ Copyright 2014 Magnolia Press Article http://dx.doi.org/10.11646/zootaxa.3753.2.9 http://zoobank.org/urn:lsid:zoobank.org:pub:af18449f-e216-44c3-a7b0-0e9dee44b4c7
Zootaxa 3753 (2): 196 200 www.mapress.com/zootaxa/ Copyright 2014 Magnolia Press Article http://dx.doi.org/10.11646/zootaxa.3753.2.9 http://zoobank.org/urn:lsid:zoobank.org:pub:af18449f-e216-44c3-a7b0-0e9dee44b4c7
REVISTA NORDESTINA DE BIOLOGIA A NEW SPECIES OF ALPHEUS (CRUSTACEA, CARIDEA) FROM THE PACIFIC COAST OF COLOMBIA ABSTRACT
 Revta. nordest. Biol., 6(1): 61-65. REVISTA NORDESTINA DE BIOLOGIA 4f V V 15.V.1988 A NEW SPECIES OF ALPHEUS (CRUSTACEA, CARIDEA) FROM THE PACIFIC COAST OF COLOMBIA M. L. Christoffersen and G.E. Ramos
Revta. nordest. Biol., 6(1): 61-65. REVISTA NORDESTINA DE BIOLOGIA 4f V V 15.V.1988 A NEW SPECIES OF ALPHEUS (CRUSTACEA, CARIDEA) FROM THE PACIFIC COAST OF COLOMBIA M. L. Christoffersen and G.E. Ramos
ox4tates )J ieuican%usellm Groups of Lizards in the Genus Sceloporus Karyotypes of the Five Monotypic Species BY CHARLES J. COLE
 )J ieuican%usellm ox4tates PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, N. Y. I0024 NUMBER 2450 FEBRUARY II, I971 Karyotypes of the Five Monotypic Species
)J ieuican%usellm ox4tates PUBLISHED BY THE AMERICAN MUSEUM OF NATURAL HISTORY CENTRAL PARK WEST AT 79TH STREET, NEW YORK, N. Y. I0024 NUMBER 2450 FEBRUARY II, I971 Karyotypes of the Five Monotypic Species
